If you're seeing this message, it means we're having trouble loading external resources on our website.
If you're behind a web filter, please make sure that the domains *.kastatic.org and *.kasandbox.org are unblocked.
To log in and use all the features of Khan Academy, please enable JavaScript in your browser.

AP®︎/College Biology
Course: ap®︎/college biology > unit 6.
- DNA and chromatin regulation
- Regulation of transcription
- Cellular specialization (differentiation)
- Non-coding RNA (ncRNA)
- Operons and gene regulation in bacteria
- Overview: Gene regulation in bacteria
- The lac operon
- The trp operon
- Overview: Eukaryotic gene regulation
Transcription factors
- Regulation of gene expression and cell specialization
Key points:
- Transcription factors are proteins that help turn specific genes "on" or "off" by binding to nearby DNA.
- Transcription factors that are activators boost a gene's transcription. Repressors decrease transcription.
- Groups of transcription factor binding sites called enhancers and silencers can turn a gene on/off in specific parts of the body.
- Transcription factors allow cells to perform logic operations and combine different sources of information to "decide" whether to express a gene.
Introduction
Transcription: the key control point.
- If a gene is not transcribed in a cell, it can't be used to make a protein in that cell.
- If a gene does get transcribed, it is likely going to be used to make a protein (expressed). In general, the more a gene is transcribed, the more protein that will be made. Is that always the case? Not always. Sometimes, later stages of regulation can block even large quantities of mRNA from being used to make protein. For example, imagine that a gene is transcribed a lot, but the mRNA is "chopped up" (degraded) as soon as it leaves the nucleus. This would lead to very little protein getting made.
How do transcription factors work?
Binding sites, how is this different from e. coli , turning genes on in specific body parts, example: modular mouse, evolution of development, transcription factors and cellular "logic".
- Activator A is present only in skin cells
- Activator B is active only in cells receiving "divide now!" signals (growth factors) from neighbors
- Repressor C is produced when a cell's DNA is damaged
Works cited:
- Gilbert, S. F. (2000). Anatomy of the gene: Promoters and enhancers. In Developmental biology (6th ed.). Sunderland, MA: Sinauer Associates. Retrieved from https://www.ncbi.nlm.nih.gov/books/NBK10023/#_A751_ .
- Gilbert, S. F. (2000). Silencers. In Developmental biology (6th ed.). Sunderland, MA: Sinauer Associates. Retrieved from http://www.ncbi.nlm.nih.gov/books/NBK10023/#_A777_ .
- Menke, D. B., Guenther, C., and Kingsley, D. M. (2008). Dual hindlimb control elements in the Tbx4 gene and region-specific control of bone size in vertebrate limbs. Development , 135 , 2543-2553. http://dx.doi.org/10.1242/dev.017384 .
- Wray, Gregory A. (2007). The evolutionary significance of cis -regulatory mutations. Nature Reviews Genetics , 8 , 206-216. http://dx.doi.org/10.1038/nrg2063 .
- Reece, J. B., Urry, L. A., Cain, M. L., Wasserman, S. A., Minorsky, P. V., and Jackson, R. B. (2011). Combinatorial control of gene activation. In Campbell Biology (10th ed., pp. 37). San Francisco, CA: Pearson.
- Reményi, Attila, Hans R. Schöler, and Matthias Wilmanns. (2004). Combinatorial control of gene expression. Nature Structural & Molecular Biology , 11 (9), 812. http://dx.doi.org/10.1038/nsmb820 . Retrieved from http://www.nature.com/scitable/content/Combinatorial-control-of-gene-expression-16976 .
Additional references:
Want to join the conversation.
- Upvote Button navigates to signup page
- Downvote Button navigates to signup page
- Flag Button navigates to signup page


Transcription factor binding sites identification using machine learning techniques
Center or department, thesis type, awarding institution, year awarded, rights statement, data source, usage metrics.

An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
- My Bibliography
- Collections
- Citation manager
Save citation to file
Email citation, add to collections.
- Create a new collection
- Add to an existing collection
Add to My Bibliography
Your saved search, create a file for external citation management software, your rss feed.
- Search in PubMed
- Search in NLM Catalog
- Add to Search
The AP2/ERF transcription factor SmERF1L1 regulates the biosynthesis of tanshinones and phenolic acids in Salvia miltiorrhiza
Affiliations.
- 1 Laboratory of Medicinal Plant Biotechnology, College of Pharmacy, Zhejiang Chinese Medical University, Hangzhou, Zhejiang 310053, PR China; Institute of Plant Biotechnology, College of Life and Environment Sciences, Shanghai Normal University, Shanghai 200234, PR China.
- 2 Institute of Plant Biotechnology, College of Life and Environment Sciences, Shanghai Normal University, Shanghai 200234, PR China.
- 3 Laboratory of Medicinal Plant Biotechnology, College of Pharmacy, Zhejiang Chinese Medical University, Hangzhou, Zhejiang 310053, PR China.
- 4 Jiangsu Provincial Key Laboratory of Coastal Wetland Biological Resources and Environmental Protection, School of Marine and Biological Engineering, Yancheng Teachers Uninversity, Yancheng, Jiangsu Province 224051, PR China.
- 5 Laboratory of Medicinal Plant Biotechnology, College of Pharmacy, Zhejiang Chinese Medical University, Hangzhou, Zhejiang 310053, PR China; Institute of Plant Biotechnology, College of Life and Environment Sciences, Shanghai Normal University, Shanghai 200234, PR China. Electronic address: [email protected].
- PMID: 30372953
- DOI: 10.1016/j.foodchem.2018.08.119
Tanshinones and phenolic acids are two important metabolites synthesized by the traditional Chinese medicinal plant Salvia miltiorrhiza. There is increasing market demand for these compounds. Here, we isolated and functionally characterized SmERF1L1, a novel JA (Jasmonic acid)-responsive gene encoding AP2/ERF transcription factor, from Salvia miltiorrhiza. SmERF1L1 was responsive to methyl jasmonate (MJ), yeast extraction (YE), salicylic acid (SA) and ethylene treatments. Subcellular localization assay indicated that SmERF1L1 located in the nucleus. Overexpression of SmERF1L1 significantly increased tanshinones production in transgenic S. miltiorrhiza hairy roots by comprehensively upregulating tanshinone biosynthetic pathway genes, especially SmDXR. Yeast one-hybrid (Y1H) and electrophoretic mobility shift assay (EMSA) showed that SmERF1L1 binds to the GCC-box of SmDXR promoter while dual luciferase (Dual-LUC) assay showed that SmERF1L1 positively regulated the expression of SmDXR. Our study suggested that the SmERF1L1 may be a good potential target for further metabolic engineering of bioactive component biosynthesis in S. miltiorrhiza.
Keywords: AP2/ERF transcription factor; Biosynthesis; Caffeic acid (PubChem CID: 689043); Cryptotanshinone (PubChem CID: 160254); Dihydrotanshinone (PubChem CID: 11425923); Phenolic acids; Rosmarinic acid (PubChem CID: 5315615); Salvia miltiorrhiza; Salvianolic acid A (PubChem CID: 5281793); Salvianolic acid B (PubChem CID: 11629084); Tanshinone I (PubChem CID: 114917); Tanshinone IIA (PubChem CID: 164676); Tanshinones.
Copyright © 2018 Elsevier Ltd. All rights reserved.
PubMed Disclaimer
Similar articles
- MAPKK2/4/5/7-MAPK3-JAZs modulate phenolic acid biosynthesis in Salvia miltiorrhiza. Xie Y, Ding M, Yin X, Wang G, Zhang B, Chen L, Ma P, Dong J. Xie Y, et al. Phytochemistry. 2022 Jul;199:113177. doi: 10.1016/j.phytochem.2022.113177. Epub 2022 Mar 28. Phytochemistry. 2022. PMID: 35358599 Review.
- The biosynthesis of phenolic acids is positively regulated by the JA-responsive transcription factor ERF115 in Salvia miltiorrhiza. Sun M, Shi M, Wang Y, Huang Q, Yuan T, Wang Q, Wang C, Zhou W, Kai G. Sun M, et al. J Exp Bot. 2019 Jan 1;70(1):243-254. doi: 10.1093/jxb/ery349. J Exp Bot. 2019. PMID: 30299490
- Overexpression of SmbHLH148 induced biosynthesis of tanshinones as well as phenolic acids in Salvia miltiorrhiza hairy roots. Xing B, Liang L, Liu L, Hou Z, Yang D, Yan K, Zhang X, Liang Z. Xing B, et al. Plant Cell Rep. 2018 Dec;37(12):1681-1692. doi: 10.1007/s00299-018-2339-9. Epub 2018 Sep 18. Plant Cell Rep. 2018. PMID: 30229287
- Comprehensive transcriptome profiling of Salvia miltiorrhiza for discovery of genes associated with the biosynthesis of tanshinones and phenolic acids. Zhou W, Huang Q, Wu X, Zhou Z, Ding M, Shi M, Huang F, Li S, Wang Y, Kai G. Zhou W, et al. Sci Rep. 2017 Sep 5;7(1):10554. doi: 10.1038/s41598-017-10215-2. Sci Rep. 2017. PMID: 28874707 Free PMC article.
- The Biosynthetic Pathways of Tanshinones and Phenolic Acids in Salvia miltiorrhiza. Ma XH, Ma Y, Tang JF, He YL, Liu YC, Ma XJ, Shen Y, Cui GH, Lin HX, Rong QX, Guo J, Huang LQ. Ma XH, et al. Molecules. 2015 Sep 8;20(9):16235-54. doi: 10.3390/molecules200916235. Molecules. 2015. PMID: 26370949 Free PMC article. Review.
- SmEIL1 transcription factor inhibits tanshinone accumulation in response to ethylene signaling in Salvia miltiorrhiza . Li X, Xu M, Zhou K, Hao S, Li L, Wang L, Zhou W, Kai G. Li X, et al. Front Plant Sci. 2024 Apr 2;15:1356922. doi: 10.3389/fpls.2024.1356922. eCollection 2024. Front Plant Sci. 2024. PMID: 38628367 Free PMC article.
- Overexpression of AtMYB2 Promotes Tolerance to Salt Stress and Accumulations of Tanshinones and Phenolic Acid in Salvia miltiorrhiza . Li T, Zhang S, Li Y, Zhang L, Song W, Chen C. Li T, et al. Int J Mol Sci. 2024 Apr 8;25(7):4111. doi: 10.3390/ijms25074111. Int J Mol Sci. 2024. PMID: 38612919 Free PMC article.
- DoAP2/ERF89 activated the terpene synthase gene DoPAES in Dendrobium officinale and participated in the synthesis of β- patchoulene. Li D, Liu L, Li X, Wei G, Cai Y, Sun X, Fan H. Li D, et al. PeerJ. 2024 Jan 18;12:e16760. doi: 10.7717/peerj.16760. eCollection 2024. PeerJ. 2024. PMID: 38250724 Free PMC article.
- The SmERF1b-like regulates tanshinone biosynthesis in Salvia miltiorrhiza hairy root. Li D, Liu Y, Chen G, Yan Y, Bai Z. Li D, et al. AoB Plants. 2023 Dec 8;16(1):plad086. doi: 10.1093/aobpla/plad086. eCollection 2024 Jan. AoB Plants. 2023. PMID: 38249522 Free PMC article.
- Adipose tissue inflammation linked to obesity: A review of current understanding, therapies and relevance of phyto-therapeutics. Aruwa CE, Sabiu S. Aruwa CE, et al. Heliyon. 2023 Dec 2;10(1):e23114. doi: 10.1016/j.heliyon.2023.e23114. eCollection 2024 Jan 15. Heliyon. 2023. PMID: 38163110 Free PMC article. Review.
- Search in MeSH
Related information
- PubChem Compound (MeSH Keyword)
LinkOut - more resources
Full text sources.
- Elsevier Science
Research Materials
- NCI CPTC Antibody Characterization Program
- Citation Manager
NCBI Literature Resources
MeSH PMC Bookshelf Disclaimer
The PubMed wordmark and PubMed logo are registered trademarks of the U.S. Department of Health and Human Services (HHS). Unauthorized use of these marks is strictly prohibited.
- Open access
- Published: 21 June 2024
Characterization of YABBY transcription factors in Osmanthus fragrans and functional analysis of OfYABBY12 in floral scent formation and leaf morphology
- Tingting Shi 1 , 2 ,
- Ling Zhou 1 , 2 ,
- Yunfang Ye 1 , 2 ,
- Xiulian Yang 1 , 2 ,
- Lianggui Wang 1 , 2 &
- Yuanzheng Yue 1 , 2
BMC Plant Biology volume 24 , Article number: 589 ( 2024 ) Cite this article
33 Accesses
Metrics details
The plant-specific YABBY transcription factor family plays important roles in plant growth and development, particularly leaf growth, floral organ formation, and secondary metabolite synthesis.
Here, we identified a total of 13 OfYABBY genes from the Osmanthus fragrans genome. These 13 OfYABBY genes were divided into five subfamilies through phylogenetic analysis, and genes in the same subfamily showed similar gene structures and conserved protein motifs. Gene duplication promoted the expansion of the OfYABBY family in O . fragrans . Tissue-specific expression analysis showed that the OfYABBY family was mainly expressed in O . fragrans leaves and floral organs. To better understand the role of OfYABBY genes in plant growth and development, OfYABBY12 was selected for heterologous stable overexpression in tobacco, and OfYABBY12 -overexpressing tobacco leaves released significantly fewer volatile organic compounds than wild-type tobacco leaves. Overexpression of OfYABBY12 led to the downregulation of NtCCD1/4 and decreased β-ionone biosynthesis. Correspondingly, a dual-luciferase assay showed that OfYABBY12 negatively regulated the expression of OfCCD4 , which promotes β-ionone synthesis. Furthermore, tobacco leaves overexpressing OfYABBY12 were curled and wrinkled and had significantly reduced leaf thickness and leaf inclusions and significantly extended flower pistils (styles).
Overall, the results suggest that the OfYABBY gene family may influence the biosynthesis of the floral scent (especially β-ionone) in O . fragrans and may regulate leaf morphogenesis and lateral organs.
Peer Review reports
Introduction
Osmanthus fragrans is an important ornamental evergreen tree or shrub that is widely cultivated for its pleasing floral scent. It is commonly used in the landscaping and fragrance industries, where it has significant ornamental and economic value [ 1 , 2 ]. Many O . fragrans cultivars have been cultivated in China for more than 2,500 years. These cultivars have significant differences in floral scent, floral color, leaf shape, and leaf color [ 3 , 4 , 5 ]. Previous studies have shown that the YABBY gene family not only participates in the formation and development of plant leaves and floral organs but also affects the biosynthesis of secondary metabolites, such as terpene floral metabolites [ 6 , 7 , 8 ]. In this study, it was hypothesized that the phenotypic difference in the leaves, flowers, and floral scents of different O . fragrans cultivars may be regulated by the O . fragrans YABBY family.
The YABBY family is a class of plant-specific transcription factors [ 9 ]. Due to the roles these transcription factors play in many of the biological processes of plants, the YABBY family has attracted great interest from researchers. The YABBY proteins contain two conserved domains, an N-terminal C 2 C 2 zinc finger domain and a C-terminal YABBY domain. In Arabidopsis thaliana , Solanum lycopersicum , and other dicotyledonous plants, the YABBY transcription factors are classified into five subfamilies, namely CRABS CLAW (CRC), FILAMENTOUS FLOWER (FIL)/YABBY3 (YAB3), INNER NO OUTER (INO), YABBY2 (YAB2), and YABBY5 (YAB5). However, in Oryza sativa , Zea mays , and other monocotyledons, contain only four subfamilies and lack the YAB5 subfamily [ 9 ]. It has been indicated that the YABBY transcription factors in monocotyledonous and dicotyledonous plants have undergone functional divergence, and four gene duplication events have occurred in the YABBY family, leading to genes with innovative or redundant functions [ 10 ].
Some studies have shown that YABBY transcription factors are related to abaxial axis cell differentiation in lateral organs, thereby affecting leaf growth [ 11 , 12 ] or the development of floral organs [ 13 , 14 ] and fruit (seeds) [ 15 , 16 ]. The YABBY family also participates in the biosynthesis of primary and secondary plant metabolites [ 8 , 17 , 18 ] and in response to biotic and abiotic stress [ 19 , 20 , 21 ]. In A. thaliana , AtYAB2 , AtFIL , AtYAB3 , and AtYAB5 are expressed in the cotyledons, leaves, petals, stamens, and carpels and participate in regulating bud and leaf development and the formation of leaf polarity and floral organs [ 6 , 7 ]. However, AtCRC is only expressed in nectaries and carpels, regulating the abaxial development of these structures [ 22 ]. Similarly, AtINO is only expressed in the abaxial axis of the outer integument and mainly regulates ovule development, thereby affecting seed growth and development [ 23 , 24 ]. In S. lycopersicum , SlYABBY1 regulates leaf and flower sizes [ 25 ], SlYABBY2 regulates pericarp development and fruit maturation [ 26 ], and SlCRCa regulates flower and fruit size [ 27 ]. In Chrysanthemum morifolium , CmDRP (FlL/YAB3 subfamily) can regulate gibberellin biosynthesis to affect chrysanthemum plant height [ 28 ]. In addition, one study has shown that YABBY genes have dual functions, as they can act as both an activator and a repressor of secondary metabolites [ 29 ]. For example, MsYABBY5 of Mentha spicata negatively regulates the synthesis of monoterpenes [ 8 ], whereas in Artemisia annua , AaYABBY5 positively regulates the biosynthesis of artemisinin [ 17 ] and flavonoids [ 18 ]. Hence, YABBY transcription factors show tissue specificity in plants and play a variety of roles in plant growth and development.
Here, a comprehensive analysis of the O . fragrans YABBY gene family was conducted based on whole genome data, and OfYABBY12 was stably overexpressed in tobacco to investigate whether OfYABBY12 affects the growth and development of leaves and floral organs, as well as the synthesis of volatile organic compounds (VOCs). These results help to understand the OfYABBY family in O . fragrans and provide insight into the function of the OfYABBY12 gene. The findings have important significance for further study of the functions of the OfYABBY family in O . fragrans.
Materials and methods
Plant materials.
The experimental material was three-year-old O. fragrans ‘Rixiang Gui’ cuttings obtained from our O . fragrans germplasm resource bank (Jiangsu, China). Various tissues, including roots, stems, young leaves (from current-year branchlets), and mature leaves (from non-current-year branchlets) were sampled on March 26, 2022, and flowers at the full blooming stage were sampled on October 9, 2022. Nicotiana benthamiana and Nicotiana tabacum ‘K326’ seeds were saved by our group and planted in the greenhouse of Nanjing Forestry University under the following growth conditions: a 16/8 h light/dark cycle, a light intensity of 110 µmol photons m − 2 s − 1 white light, a temperature of 25 ± 2 °C, and 60–70% relative humidity.
Identification of O. fragrans YABBY transcription factors and protein physicochemical analysis
The whole genome sequence of O . fragrans was obtained from the O . fragrans ‘Rixiang Gui’ genomic database [ 30 ]. HMMER software (v3.0) was then used to screen for family members in the Pfam YABBY (PF04690) domain [ 31 ]. Three online tools, Batch CD-search ( http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi ), Pfam ( http://pfam.xfam.org/search#searchBatchBlock ), and SMART ( http://smart.embl.de/smart/batch.pl ), were used to validate the YABBY domains. In addition, ExPASy ( https://web.expasy.org/compute_pi/ ) was employed to predict the molecular weight and isoelectric point of the identified YABBY protein sequences.
Gene structure, conserved motif, and cis-acting element analysis
In this study, DNAMAN software (v6.0.40) was used for multi-sequence alignment and the analysis of the conserved protein domains of YABBY amino acid sequences in O . fragrans . In addition, MEME online software ( http://meme-suite.org/ ) was used to analyze the conserved motifs of the OfYABBY protein sequences. The gene sequences 2,000 bp upstream of the start codon were obtained from the O . fragrans genome sequences, and the plantCARE database ( http://bioinformatics.psb.ugent.be/webtools/plantcare/html/ ) was used to retrieve the obtained sequences to analyze the potential cis-acting elements in the promoter. Finally, TBtools software (v1.120) [ 32 ] was used to visualize the gene structure, conserved motifs, and cis-acting elements of OfYABBYs.
Phylogenetic, chromosome localization, and gene duplication analysis
Genomic data were obtained from different databases, including A . thaliana ( https://www.arabidopsis.org/ ), S . lycopersicum ( https://solgenomics.net/ ), O. sativa ( http://rice.uga.edu/ ), and Z. mays ( https://maize-pangenome.gramene.org/ ). Furthermore, the protein sequences of YABBYs from Vitis vinifera [ 15 ], Camellia sinensis [ 33 ], Mimulus lewisii [ 34 ], M . spicata [ 8 ], Chimonanthus praecox [ 35 ], and A . annua [ 17 ] were obtained from relevant references.
Clustal X2.1 was used for the sequence alignment of the YABBY protein sequences of O . fragrans and other plants. In addition, MEGA X software was utilized to construct phylogenetic trees using the maximum likelihood method (Bootstrap = 1000) [ 36 ], and Fig Tree software (v1.4.3) was employed for visualization.
The chromosome locations of the OfYABBY genes were obtained based on the O . fragrans genome GFF3 file, and TBtools was used for visualization. Multiple Collinearity Scan Toolkits (MCScanX) and Ka/Ks calculators (NG) were used to further analyze the gene duplication events of the OfYABBY genes.
RNA extraction and quantitative reverse-transcription polymerase chain reaction (qRT-PCR) analyses
The RNAprep Pure Plant Kit (Tiangen, Beijing, China) was used to obtain the total RNA of plant materials, and One-Step gDNA Removal and cDNA Synthesis Super Mix (Transgen, Beijing, China) were used for reverse transcription to obtain complementary DNA (cDNA).
Primer Premier 5.0 software was used to design the qRT-PCR primers (Table S1 ). The ABI StepOne Plus PCR system and SYBR® Premix Ex Taq™ II Kit (Takara, Dalian, China) were used to measure gene expression levels. In addition, 10-fold diluted cDNA was used as a template, and OfACTIN was used as an internal reference gene [ 37 ]. The 2 −ΔΔCT method [ 38 ] was employed to calculate the relative expression level. Biological triplicates were set up for each qRT-PCR reaction, and technical triplicates were set up for each biological triplicate.
Subcellular localization and transactivation assay
The cDNAs of OfYABBY3 , OfYABBY5 , OfYABBY7 , OfYABBY8 , OfYABBY12 , and OfYABBY13 were cloned (primers shown in Table S2 ) and inserted into pCAMBIA1300 vectors to build the pCAMBIA1300-GFP-OfYABBY constructs. These constructed plasmids were transformed into N . benthamiana leaves by Agrobacterium tumefaciens strain GV3101 (Weidi Biotechnology, Shanghai, China). After 48 h, a Zeiss LSM 710 confocal microscope (Zeiss, Oberkochen, Germany) was used to observe fluorescence signals.
The cDNAs of OfYABBY3 , OfYABBY5 , OfYABBY7 , OfYABBY8 , OfYABBY12 , and OfYABBY13 were cloned into the pGBKT7 vector (primers shown in Table S2 ). These recombination vectors and the pGBKT7 empty vector were transformed into Saccharomyces cerevisiae (AH109). The suspended yeast was then diluted to different concentrations, and 5 µL dilutions were plated on SD-Trp, SD-Trp/Ade, and SD-Trp/Ade (X-α-gal) culture media. These were cultured at 30 °C in the dark for 3 d, and the growth status was then observed.
Transformation of OfYABBY12 in N. tabacum
A. tumefaciens cells carrying the pCAMBIA1300-OfYABBY12 plasmid were cultured overnight, and the culture products were collected when the OD600 reached 0.4–0.5. A solution of 10 mM MgCl 2 , 10 mM MES, and 150 mM acetosyringone was used to resuspend the Agrobacterium cultures for transformation. In addition, tobacco ( N . tabacum ‘K326’) seeds were sterilized and sown on MS medium, and when grown to three to four leaves, sterile tobacco leaves were cut into 0.5 cm × 0.5 cm cubes (that is, explants) and infiltrated with Agrobacterium cultures for 10 min. Subsequently, after 3 d of co-cultivation in the dark on the co-culture medium (MS + 2.25 mg/L 6-BA + 0.3 mg/L NAA), the explants were transferred to selective medium (MS + 2.25 mg/L 6-BA + 0.3 mg/L NAA + 400 mg/L cefotaxime + 100 mg/L kanamycin). The culture was kept at 25 °C with a 16/8 h light/dark cycle with the medium changed every 15 d. When the callus and resistant shoots emerged, the explants were transferred to germination medium (MS + 0.1 mg/L 6-BA + 0.01 mg/L NAA + 400 mg/L cefotaxime + 100 mg/L kanamycin). When the shoots grew to around 2 cm in length, the explants were transferred to rooting medium (MS + 400 mg/L cefotaxime + 100 mg/L kanamycin). Finally, after the roots were developed and mature, the plantlets were removed from the medium and planted in sterile soil.
After one month of growth, a Plant Rapid Genome Extraction kit (Tsingke, Wuhan, China) was used for a positive test of pCAMBIA1300-OfYABBY12 transgenic and wild-type (WT) tobacco leaves. Positive plants and WT plants were used for semi-quantitative validation, and transgenic lines with high levels of expression were selected for phenotype observation, scanning electron microscopy, gas chromatography–mass spectrometry (GC–MS), and RNA sequencing (RNA-Seq).
GC–MS volatile analysis
Headspace solid-phase microextraction (HS-SPME) was used to collect VOCs released from tobacco leaves or flowers. The samples of fresh leaves or flowers were ground in liquid nitrogen, 1 g of leaf or flower powder was placed in a 10-mL SPME vial with 3 mL NaCl saturated solution, and 1 µL of 5000-fold diluted ethyl caprate was added as the internal standard. Each vial was placed at room temperature (25 ± 2 °C) for 30 min and then extracted at 55 °C for 30 min using an extraction head composed of 65 µM polydimethylsiloxane (PDMS)/divinylbenzene (DVB) fiber (Supelco Co., Bellefonte, PA, USA) for GC–MS detection. GC–MS sequencing was conducted according to a previously described protocol [ 30 ].
Scanning electron microscopy
Fresh tobacco leaf samples were fixed in formalin–acetic acid–ethyl alcohol (FAA) at 4 °C for 48 h. The samples were then dehydrated sequentially in a gradient ethanol series, and after dehydration, the samples were vacuum-dried and sprayed with a gold film layer. Finally, the samples were placed under an environmental scanning electron microscope (Quanta 200, FEI Inc, Netherlands) for observation and photographs.
RNA-Seq analysis
Total RNA was extracted from mature WT and OfYABBY12 -overexpressing ( OfYABBY12-OE ) tobacco leaves using an RNAprep Pure Plant Kit (Tiangen, Beijing, China). Each sample had three biological replicates. RNA-Seq was performed by the Guangzhou Gene Denovo Biotechnology Co. (Guangzhou, China) using the Illumina HiSeq2500 platform. All sequencing data were uploaded to the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) database ( https://www.ncbi.nlm.nih.gov/sra/ ) under the accession number SRP450701.
Dual-luciferase transient expression assay
The coding sequence (CDS) of OfYABBY12 was amplified via PCR using gene-specific primers (Table S2 ) and connected to the pGreenII 62-SK vector to generate an effector, which was transformed into the A . tumefaciens strain GV3101 (pSoup). Then, the A . tumefaciens cells carrying the effector and reporter (pGreenII 0800-LUC-OfCCD4, preserved by our group) plasmids were mixed in a 4:3 ratio (v/v) and injected into N. benthamiana leaves. After 48 h, the luciferase (LUC) signal was detected using BERTHOLD NightSHADE LB 985 (Berthold, Bad Wildbad, Germany), and LUC and renilla luciferase (REN) were detected using a Dual-Luciferase Reporter Assay System kit (Yeasen Biotechnology, Shanghai, China).
Statistical analysis
One-way analysis of variance (ANOVA), Duncan’s multiple range test, and Student’s t-test were performed using IBM SPSS Statistics (version 20.0). A p -value smaller than 0.05 was considered significant. A minimum of three biological replicates were utilized for all experiments. Bar graphs were created using Origin Pro 2018. Heat maps were generated using TBtools software (version 1.120). Principal component analysis (PCA) and orthogonal partial least squares discrimination analysis (OPLS-DA) of VOCs were performed using SIMCA-13.0 software (version 13.0.3).
Identification and classification of YABBY gene family members in O. fragrans
In this study, HMMER was used to identify 19 candidate YABBY genes in the O . fragrans whole genome database, and six sequences without intact structural domains were removed using NCBI-CDD, Pfam, and SMART. A total of 13 YABBY genes were identified in the O . fragrans genome. The OfYABBY genes were named based on their position on chromosomes as OfYABBY1–OfYABBY13 , and these 13 genes were located on O . fragrans chromosomes 6, 10, 11, 16, 21, and 23 (Fig. 1 A). The length of OfYABBY proteins ranged from 149 aa (OfYABBY13) to 247 aa (OfYABBY2), and the protein molecular weight ranged from 16.69 kDa (OfbYABBY13) to 27.39 kDa (OfbYABBY2). The theoretical isoelectric point (pI) was 6.18–9.3, and OfYABBY5, OfYABBY8, OfYABBY9, and OfYABBY10 were acidic proteins (pI < 7.00), while the remaining proteins were basic (Table S3 ).
The sequence alignment results of the 13 OfYABBY proteins revealed that all OfYABBY proteins contained two highly conserved protein domains, of which one was a C 2 C 2 zinc finger domain in the N terminal, and one was a YABBY domain in the C terminal (Fig. S1 A). In addition, the 13 OfYABBY genes contained four to seven introns, of which most OfYABBY genes contained six introns. OfYABBY5 and OfYABBY 8 contained the greatest number of introns at seven. Overall, OfYABBY genes that clustered together shared identical or similar exon/intron structures. The conserved motifs of the OfYABBY family were analyzed (Table S4 ). All members contained at least two motifs, motifs 1 and 2, and genes that clustered together had similar motifs (Fig. S1 B). These genes may have similar functions.
Interactions between cis- and trans-acting factors can regulate gene expression. The promoters and their upstream cis-acting elements were analyzed to predict their corresponding gene functions. In the O . fragrans YABBY gene family, every gene promoter contained four to 10 cis-acting elements (Fig. S2 ). Most were considered involved in light response, including GT1-motif, Box 4, TCT-motif, G-Box, AE-box, GA-motif, ATCT-motif, GATA-motif, chs-CMA1a, 3-AF1 binding, I-box, AE-box, TCCC-motif, MRE, and ACE. The responses to gibberellin, abscisic acid, salicylic acid, auxin, and methyl jasmonate included TATC-box, P-box, GARE-motif, ABRE, TCA-element, TGA-box, TGA-element, TGACG-motif, and CGTCA-motif. LTR, MBS, ARE, and TC-rich repeats might be involved in responding to drought, anaerobic, and low temperature stress. In addition, cis-acting elements such as CAT-box and HD-Zip 1 were involved in the developmental regulation of plant tissues and organs. This means that OfYABBY s may be involved in the responses to many plant hormones and environmental stress and play a role in plant growth and development.
Phylogenetic analysis and gene duplication
To understand the phylogenetic relationship between various YABBYs, 56 YABBY proteins from dicotyledonous plants ( O . fragrans , A . thaliana , S . lycopersicum , and V . vinifera [ 15 ]) and monocotyledonous plants ( O . sativa and Z . mays ) were used to construct a phylogenetic tree (Fig. 1 B). Based on the classification of A. thaliana YABBY proteins [ 39 ], the 13 OfYABBYs were divided into five subfamilies, namely FIL/YAB3, YAB5, YAB2, CRC, and INO. The FIL/YAB3 subfamily included five OfYABBY proteins, and the YAB2 subfamily included four OfYABBY proteins. In addition, the INO subfamily included two YABBY proteins, and the CRC and YAB5 subfamilies each included one OfYABBY (Table S3 ). As predicted, phylogenetic analysis showed that OfYABBYs had the closest phylogenetic relationship to YABBYs from dicotyledonous plants. The YABBYs of dicotyledonous plants were all clustered together. Moreover, the YABBYs of monocotyledonous plants, such as rice ( O . sativa ), formed a parallel branch, consistent with previous reports [ 9 ]. This shows that YABBY genes may function in the differentiation of monocotyledonous and dicotyledonous plants and that they are conserved in dicotyledonous plants.
In addition, phylogenetic analysis of O . fragrans YABBY protein sequences was conducted based on YABBYs in M . lewisii , C . sinensis , C . praecox , A . annua , and M . spicata . This is because their functions have been sufficiently discussed, such as MlYAB1 / 2 / 3 / 5 , CsFILa / b , and CsYAB2 , which are involved in leaf development regulation [ 33 , 34 ], and CpFIL , CpCRC , CpYABBY2 , CpYABBY5-1 / -2 , AaYABBY5 and MsYABBY5 , which are involved in secondary metabolite biosynthesis [ 8 , 17 , 18 , 35 ]. Based on the results (Fig. S3 ), it was deduced that the functions of OfYABBY1 / 2 / 3 / 4 / 5 / 6 / 7 / 8 / 12 / 13 genes of the FIL/YAB3, YAB2, and YAB5 subfamilies in O. fragrans may be related to leaf development or secondary metabolite synthesis. In addition, as shown in Fig. 1 B, the FIL/YAB3 and YAB2 subfamilies contained more OfYABBY genes than other subfamilies, exhibiting significant gene duplication. However, because the number of O . fragrans YABBY family members was low, gene collinearity analysis did not find tandem repeat OfYABBY genes, but three segmental duplication of OfYABBY genes ( OfYABBY1/OfYABBY12 , OfYABBY2/OfYABBY13 , and OfYABBY4/OfYABBY7 ) were detected (Fig. 1 C).
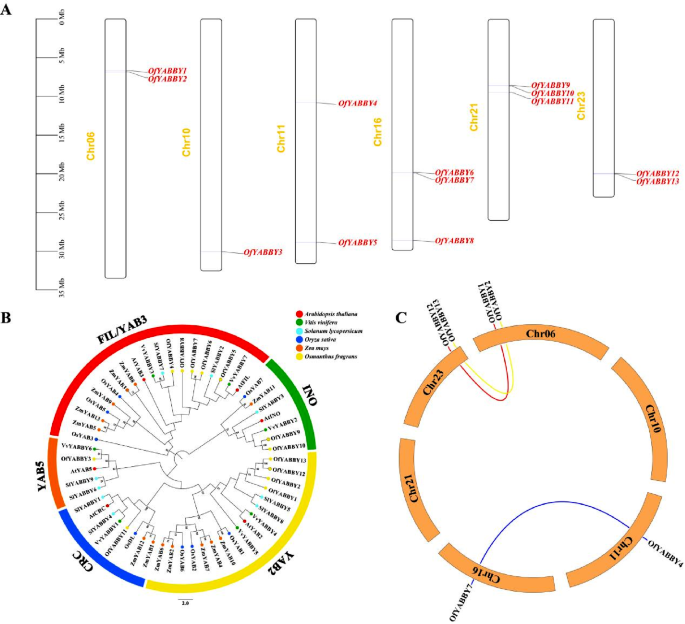
Chromosomal distribution, phylogenetic relationships, and collinearity analysis of Osmanthus fragrans YABBY genes. ( A ) Chromosome locations of the OfYABBY genes. ( B ) Phylogenetic analysis of YABBYs from O . fragrans , Arabidopsis thaliana , Vitis vinifera , Solanum lycopersicum , Oryza sativa and Zea mays . Numbers at the nodes indicate bootstrap values; values lower than 50% are not shown. The sequences of the YABBYs used for phylogenetic relationship analysis are listed in Table S5 . ( C ) Collinearity analysis for all OfYABBY genes
Expression patterns of OfYABBYs
Transcriptome data were obtained to analyze the expression patterns of OfYABBY s in different O . fragrans cultivars [ 40 ] and different flowering stages (bud-pedicel stage, bud-eye stage, primary blooming stage, full blooming stage, and flower fading stage) (unpublished, Table S6 ). The results showed that two OfYABBY genes in the INO subfamily ( OfYABBY9 and OfYABBY10 ) and OfYABBY11 in the CRC subfamily had limited expression or were not expressed in floral tissues. Previous studies have reported that INO and CRC subfamily genes are only expressed in reproductive organs, and it is hypothesized in this study that OfYABBY s in the CRC and INO subfamilies may not participate in the formation of floral organs. Unlike OfYABBY9 , OfYABBY10 , and OfYABBY11 , OfYABBY genes in the FIL/YAB3, YAB2, and YAB5 subfamilies, except from OfYABBY4 and OfYABBY8 , were highly expressed in floral tissues, especially OfYABBY12 and OfYABBY13 (Fig. 2 A).
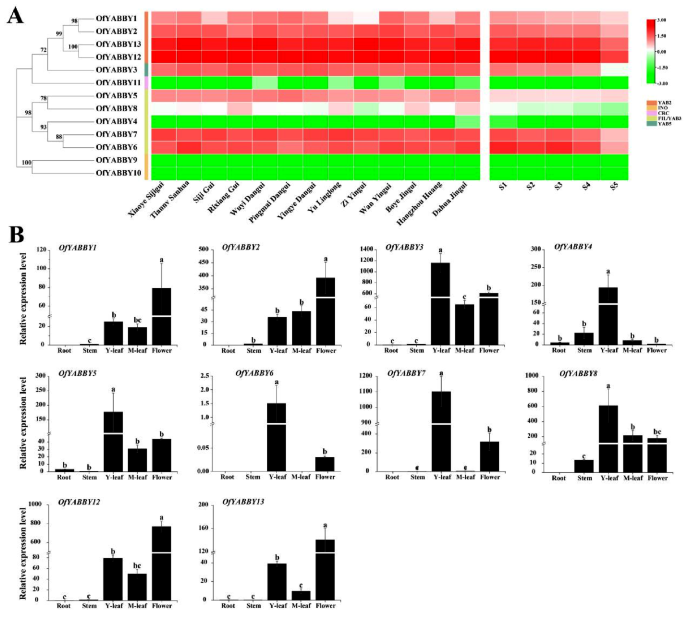
Expression analysis of OfYABBY genes. ( A ) Differential expression profiles of OfYABBY genes in 13 cultivars at the full blooming stage, and five flowering stages including the bud-pedicel stage ( S1 ), bud-eye stage ( S2 ), primary blooming stage ( S3 ), full blooming stage ( S4 ), and flower fading stage ( S5 ) of Osmanthus fragrans ‘Rixiang Gui’ based on RNA sequencing data. ( B ) Expression patterns of OfYABBY genes were confirmed in roots, stems, leaves (young and mature leaves), and flowers (full blooming stage) using quantitative reverse-transcription polymerase chain reaction (qRT-PCR) analysis. Error bars indicate the standard deviations of three biological replicates. Different letters indicate a significant difference ( p < 0.05) as determined by analysis of variance (ANOVA), which is based on Duncan’s multiple range test
This study further conducted qRT-PCR analysis of the expression levels of 10 OfYABBY genes from the FIL/YAB3, YAB2, and YAB5 subfamilies in the roots, stems, leaves (young and mature leaves), and floral tissues. The expression patterns of various OfYABBY s in different tissues showed significant differences (Fig. 2 B). Overall, these 10 OfYABBY genes were slightly or not expressed in roots and stems. Among them, OfYABBY4/5/6/7/8 from the FIL/YAB3 subfamily exhibited preferential expression in young leaves, and OfYABBY3 from the YAB5 subfamily had low expression in roots, stems, and mature leaves, and highest expression in young leaves, followed by flowers. Notably, OfYABBY1 /2 /12/13 exhibited similar expression patterns, with significantly higher expression in flowers than in other tissues.
Subcellular localization and transcriptional activation activity of OfYABBYs
To examine the potential function of OfYABBY genes in transcriptional regulation, this study selected six OfYABBY genes from the FIL/YAB3, YAB2, and YAB5 subfamilies for subcellular localization analysis based on the bioinformatics analysis. Laser confocal microscopy revealed that the green fluorescent protein (GFP) fluorescence signals of 35 S::GFP-OfYABBY3/5/7/8/12/13 fusion proteins were detected mainly in the cell nuclei (Fig. 3 ). In addition, this work further cloned these six OfYABBY genes into the yeast expression vector pGBKT7 and found that yeast strains carrying the pGBKT7 empty vector (negative control) and the pGBKT7-OfYABBY vectors exhibited good growth on SD/-Trp culture medium. Only pGBKT7-OfYABBY5/8-transformed yeast could grow on SD/-Trp-Ade culture medium, and these strains showed normal growth and blue colonies on X-α-gal containing SD/-Trp-Ade culture medium (Fig. 4 ). This indicates that OfYABBY5/8 shows transcriptional activity in yeast, while OfYABBY3/7/12/13 does not show transcriptional activity.
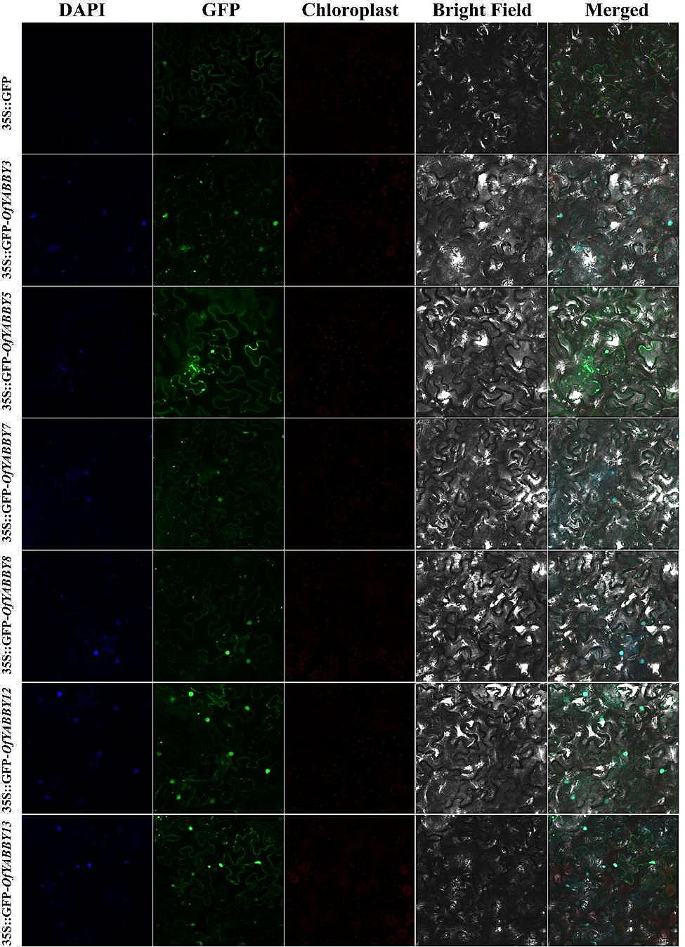
Subcellular localization of selected OfYABBY proteins. The OfYABBY proteins fused with green fluorescent protein (GFP) were transiently expressed in tobacco leaf cells to observe subcellular localization through laser confocal microscopy. GFP fluorescence is shown in green and the nucleus is blue with 4’,6-diamidino-2-phenylindole (DAPI) staining
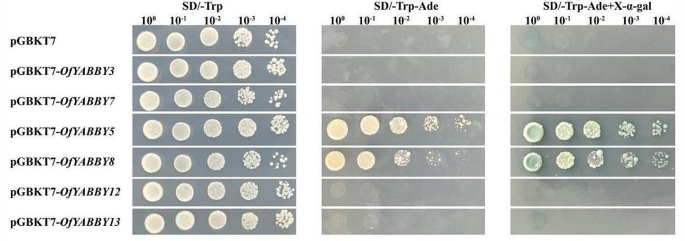
Transcriptional activation analysis of selected OfYABBY proteins. The transformed yeast cells containing pGBKT7 and pGBKT7-OfYABBYs were spread on SD/-Trp, SD/-Trp-Ade, and SD/-Trp-Ade + X-α-gal media
Phenotypes of transgenic N. tabacum overexpressing OfYABBY12
Plant YABBY transcription factors participate in the development of plant leaves and floral organs, as well as the biosynthesis of secondary metabolites, such as floral VOCs. Therefore, this study selected OfYABBY12 , significantly more highly expressed in O. fragrans flowers than in other organs, for heterologous overexpression in tobacco. This allowed the further examination of the potential function of OfYABBY s. Positive transgenic tobacco plants were selected using semi-quantitative RT-PCR, and 10 transgenic lines were obtained (Fig. S4 ). Three lines with higher expression levels (lines 2, 5, and 12) were selected for subsequent analysis.
The phenotypic observation of tobacco leaves showed that, compared with the WT, the most intuitive phenotypic difference was that wrinkles appeared at the early stages of development in OfYABBY12-OE tobacco leaves and became more severe (Fig. 5 A). Compared with WT tobacco, leaves in the overexpressing lines exhibited significant downward curvature and wrinkling (Fig. 5 B), indicating that OfYABBY12 participated in the establishment of abaxial polarity in tobacco leaves. Scanning electron microscopy images of tobacco leaves (Fig. 5 C) showed that the cross-sections of leaves from overexpression lines were significantly thinner, while the palisade and spongy mesophyll tissues were denser compared with WT tobacco (panels a–d). The leaf thickness of OfYABBY12-OE in lines 2, 5, and 12 decreased by 33, 33, and 57%, respectively, compared with the WT (Fig. S6 ). It was also observed that WT tobacco leaves were filled with granular inclusions, while there were significantly fewer inclusions in overexpressing lines (panels e–h). However, no significant difference was observed in leaf epidermal cells. Specifically, no significant differences were observed in epidermal hair (panels i–l) or stomata (panels m–p).
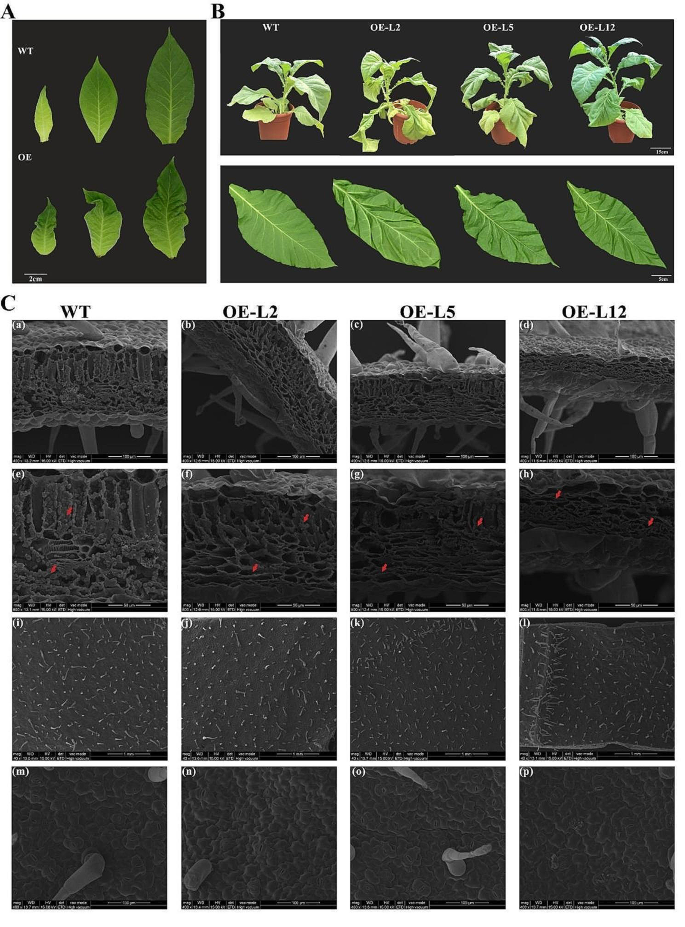
Phenotypic analysis of OfYABBY12 transgenic plants. ( A ) Leaf morphology of the wild-type (WT) and OfYABBY12-OE lines (overexpression of OfYABBY12 in tobacco (OE)) at different developmental stages. ( B ) OfYABBY12 transgenic plants produced curled leaves when compared with the WT. ( C ) Scanning electron microscopic images of tobacco leaves. (a–h) Transverse sections of tobacco leaves. (i–p) Epidermal structures of tobacco leaves. Red arrows show the inclusions in the tobacco leaves
Phenotypic observation of tobacco flowers showed that the pistil (style) of overexpressing tobacco lines was significantly extended compared with the WT, while the stamen length remained unchanged after OfYABBY12 gene expression was upregulated (Fig. S7 ).
Detection of VOCs in transgenic N. tabacum overexpressing OfYABBY12
To further determine the function of OfYABBY12 , this study assessed VOC levels in the leaves of OfYABBY12-OE tobacco plants. PCA revealed significant metabolic differences between WT and OfYABBY12-OE tobacco leaves (Fig. 6 A), and variable importance in projection (VIP) values from OPLS-DA showed that β-ionone was the key differential metabolite between WT and OfYABBY12-OE tobacco leaves (Table S7 ). In OfYABBY12-OE tobacco leaves, the β-ionone content and total VOC content were both significantly lower than in the WT leaves (Figs. 6 B and C). However, the GC–MS of the tobacco flowers showed no significant differences in VOCs released from WT and OfYABBY12-OE tobacco, and β-ionone was not detected in any of the tobacco flowers. PCA also showed that VOCs released by WT and overexpressing tobacco flowers could not be differentiated (Fig. S8 ).
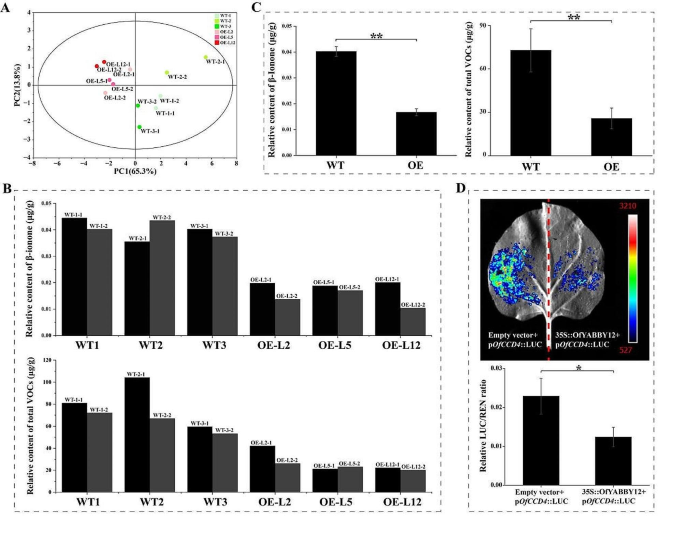
Effects of OfYABBY12 overexpression on volatile organic compound (VOC) metabolism and the structure of tobacco leaves. ( A ) Principal component analysis (PCA) of VOCs in wild-type (WT) and OfYABBY12 -overexpressing ( OfYABBY12-OE ) leaves. ( B , C ) Relative content analysis of β-ionone and total compounds of tobacco leaves. Two replicates per strain are shown in ( B ), and the means calculated from three strains with error bars reflecting standard deviations are shown in ( C ). ( D ) The dual-luciferase assay verified the relationship between OfYABBY12 and Pro- OfCCD4 . Renilla (REN) luminescence was used to normalize the luciferase (LUC) activity. The mean ± standard deviation (SD) from three replicates is shown. Asterisks indicate significant differences (Student’s t-test: * P < 0.05; ** P < 0.01)
In O . fragrans , carotenoid cleavage dioxygenase gene 4 ( CCD4 ) positively regulates β-ionone biosynthesis [ 41 , 42 ]. Thus, this study verified the regulatory relationship between OfYABBY12 and the OfCCD4 promoter using a dual-luciferase assay. Tobacco leaf cells had strong LUC signals, but the LUC/REN ratio of the experimental group (35 S::OfYABBY12 + p OfCCD4 ::LUC) was significantly lower than that of the control group (empty vector + p OfCCD4 ::LUC), indicating that OfYABBY12 negatively regulated the expression of OfCCD4 (Fig. 6 D).
Subsequently, RNA-Seq analysis was conducted on tobacco leaves to study the effects of OfYABBY12 overexpression on the transcription levels in overexpressing tobacco. A total of 3721 differentially expressed genes (DEGs, expression change > 2-fold, p < 0.05) were identified (Fig. S9 ). The Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis of DEGs showed that these DEGs mainly participate in signaling pathways and metabolism in plants. Most DEGs found in the present study were concentrated in secondary metabolite synthetic pathways, such as in the biosynthesis of terpenes, flavonoids, and phenylpropanoids (Fig. 7 A). In addition, Gene Ontology (GO) enrichment analysis of these DEGs found three GO entries in the top 20 enriched items related to terpene biosynthesis (Fig. 7 B). Analysis of the DEGs that participate in terpene biosynthesis found that most upstream genes in the metabolic pathway were upregulated in the overexpressing lines. Among downstream genes, some NtTPSs were downregulated, and some were upregulated. NtCCD1 and NtCCD4 were significantly downregulated (Fig. 7 C).

Transcriptome profiling of tobacco leaves. ( A ) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of differentially expressed genes (DEGs) in transgenic leaves. ( B ) Gene Ontology (GO) classification of unigenes among the annotated DEGs of OfYABBY12 overexpression in transgenic leaves. The bubble chart shows the enrichment of DEGs in certain pathways. ( C ) Overview of transcript changes in terpene biosynthesis pathway genes in wild-type (WT) and OfYABBY12 -overexpressing ( OfYABBY12-OE ) leaves. DXS: 1-deoxy-D-xylulose-5-phosphate synthase; DXR: 1-deoxy-D-xylulose-5-phosphate reductoisomerase; MCT: 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase; CMK: 4-diphosphocytidyl-2-C-methyl-D-erythritol kinase; MDS: 2-C-methyl-D-erythritol 2,4-cyclodiphosphate synthase; HDS: (E)-4-hydroxy-3-methylbut-2-enyl-diphosphate synthase; HDR: 4-hydroxy-3-methylbut-2-enyl diphosphate reductase; AACT: acetyl-CoA C-acetyltransferase; HMGS: 3-hydroxy-3-methylglutaryl-CoA synthase; HMGR: 3-hydroxy-3-methylglutaryl CoA reductase; MVK: mevalonate kinase; PMK: phosphomevalonate kinase; MVD: diphosphomevalonate decarboxylase; TPS: terpene synthase; CCD: carotenoid cleavage dioxygenase
Overview of the YABBY gene family in O. fragrans
The YABBY gene family has relatively low membership in different species and is a small gene family. For example, six, nine, and seven YABBY gene members have been identified in A . thaliana [ 39 ], S . lycopersicum [ 43 ], and V . vinifera [ 15 ], respectively. In this study, 13 OfYABBY genes were identified in the O . fragrans genome. This relatively high number may be the result of gene duplication events. This study did not find tandem repeats of the YABBY genes in O . fragrans . However, three segmental duplication YABBY genes were identified (Fig. 1 C), indicating that the evolution of the O . fragrans YABBY gene family may have been driven by segmental duplication. In addition, genomic studies have shown that O . fragrans underwent two whole genome duplication events compared with A . thaliana and V . vinifera [ 30 ].
OfYABBY s in O . fragrans could be divided into five subfamilies based on phylogenetic and gene structural analysis. Genes in the same subfamily showed similar motifs and intron/exon structures, suggesting that they may have similar functions. In addition, an analysis of the promoter regions of O . fragrans YABBY genes showed that photo-responsive elements were present in all members and were the cis-acting elements with the highest quantity. These genes may play an important role in photo-response and photomorphogenesis, thereby affecting the growth and development of plant leaves. Moreover, some cis-acting elements involved in the developmental regulation of plant tissues and organs were identified. Previous studies have also reported that YABBY gene family members play extremely important roles in leaf development and synthesis [ 44 , 45 , 46 , 47 ]. In addition, the cis-acting elements include some plant hormone response elements, such as abscisic acid, auxin, and methyl jasmonate. One study reported that the interactions between A . thaliana FIL and JAZ proteins (an inhibitor of the jasmonic acid pathway) affected anthocyanidin accumulation [ 48 ]. O. fragrans OfYABBY1/2/4/6/7/8/9/10/11/12/13 all contain methyl jasmonate response elements. We speculate that these OfYABBY genes may be mediated by jasmonic acid signaling to regulate the synthesis of various secondary metabolites.
The function of O . fragrans YABBY genes can be predicted to some extent based phylogenetic relationships with previously studied YABBY genes. Consistent with previous studies, the O . fragrans YABBY genes were found to have closer phylogenetic relationships with YABBYs from dicotyledon plants than with those from monocotyledon plants ( A . thaliana , S . lycopersicum , and V . vinifera ) (Fig. 1 B). As important model plants, the functions of the A . thaliana and S . lycopersicum YABBY genes have been extensively examined. These genes mainly play important roles in plant growth and development, particularly in leaf growth, fruit development, floral organ formation, and the synthesis of plant secondary metabolites [ 14 , 27 , 49 , 50 ]. In addition, subcellular localization analysis showed that 35 S::GFP-OfYABBY3/5/7/8/12/13 fusion proteins were mainly located in the nucleus, suggesting that these proteins have diverse functions in the nucleus. Transcriptional activity analysis revealed that OfYABBY5/8 exhibited transcriptional activity in yeast cells and may directly regulate downstream target genes. However, OfYABBY3/7/12/13 did not exhibit transcriptional activity in yeast cells, and thus, these proteins may require interactions with other transcription factors or specific environmental conditions to perform their regulatory functions.
OfYABBY12 plays a negative role in VOC synthesis
YABBY transcription factors have been reported to have tissue specificity in plants and to play different roles in different plant tissues [ 44 ]. In the present study, OfYABBY9 , OfYABBY10 , and OfYABBY11 were not detected or exhibited extremely low transcriptional expression in different cultivars or different flowering stages in O . fragrans floral organs. In A . thaliana , AtINO and AtCRC are only expressed in reproductive organs [ 22 , 23 ]; OfYABBY9 , OfYABBY10 , and AtINO are in the same subfamily, and OfYABBY11 and AtCRC in the same subfamily (Fig. 1 B). This suggests that O. fragrans CRC and INO subfamily genes may not participate in regulating the development of floral organs. However, the FIL/YAB3, YAB2, and YAB5 subfamily genes, including OfYABBY1/2/3/6/7/12/13 , are highly expressed in O . fragrans leaves or flowers but lowly expressed or not expressed in roots and stems. This suggests that these genes may play important roles in O . fragrans leaves and floral organs. Furthermore, studies have reported that there is redundancy in the functions of genes in the FIL/YAB3, YAB2, and YAB5 subfamilies [ 22 , 51 ], and these genes often participate in regulating leaf morphogenesis, the development and differentiation of lateral organs, and the biosynthesis of secondary metabolites [ 8 , 11 , 13 , 17 , 28 ].
Notably, comparing the transcript levels of all OfYABBY s revealed that OfYABBY12 in the YAB2 subfamily had the highest transcript level (Fig. 2 A) and showed significant differences in expression levels among O . fragrans roots, stems, young leaves, mature leaves, and floral organs (Fig. 2 B). Therefore, this study selected OfYABBY12 for further functional validation. However, a stable genetic transformation system for O . fragrans has not yet been established. Thus, OfYABBY12 was stably overexpressed in tobacco to determine whether OfYABBY12 regulates the differentiation and formation of tobacco leaves and floral organs, as well as whether it affects their volatile metabolites. In the present study, RNA-Seq analysis of WT and OfYABBY12-OE tobacco leaves found that most DEGs were enriched in metabolic pathways related to the synthesis of terpenes, flavonoids, and phenylpropanoids (Figs. 7 A and B). GC–MS results showed that there were significant differences in the VOCs between WT and overexpressing tobacco leaves. The content of β-ionone and total VOCs were significantly decreased (Figs. 6 A and C), and critical genes that participate in β-ionone synthesis, such as NtCCD1 and NtCCD4 , were significantly downregulated in overexpressing tobacco leaves (Fig. 7 C). It has also been reported that MsYABBY5 negatively regulates the synthesis of volatile terpene compounds in M . spicata [ 8 ]. β-Ionone is a critical compound for floral scent synthesis in O . fragrans [ 52 ]. Our previous study also found that OfYABBY12 in the molecular regulatory network of β-ionone synthesis showed a significant negative correlation with critical enzyme genes in its metabolic pathway (Fig. S10 ) [ 40 ]. Correspondingly, the dual-luciferase assay in the present study showed that OfYABBY12 negatively regulated the expression of OfCCD4 , which has been reported to promote β-ionone synthesis [ 41 , 42 ]. The results suggest that OfYABBY12 negatively regulates β-ionone synthesis in O . fragrans .
In addition, it was observed that the inclusions of OfYABBY12-OE tobacco leaves were significantly decreased compared with the WT under scanning electron microscopy (Fig. 5 C), but whether this is related to the reduction of VOCs needs further study. However, OfYABBY12 overexpression did not significantly affect VOCs from tobacco flowers. This suggests that OfYABBY12 overexpression affects VOCs from tobacco leaves, but not from tobacco flowers. Further studies are also needed to determine the regulatory relationship.
OfYABBY12 is involved in the development of leaves and flowers
In the present study, the overexpression of OfYABBY12 in tobacco revealed that OfYABBY12 overexpression affected the formation of the adaxial–abaxial axis in tobacco leaves, causing leaves to curl and crenations to occur. In Saccharum spontaneum , SsYABBY2 overexpression causes the abaxial side of A . thaliana leaves to curl [ 53 ]. Overexpression of C . sinensis CsFIL and CsYAB2 in A . thaliana causes leaves to curl [ 33 ]. Furthermore, one study found that the overexpression of the V . vinifera VvYABBY4 gene in S . lycopersicum caused the pistils (stigma) of transgenic S . lycopersicum to become longer [ 15 ], while the downregulation of MlYAB1 , MlYAB2 , MlYAB3 , and MlYAB5 in M . lewisii inhibited style elongation [ 34 ]. In the present study, OfYABBY12 , VvYABBY4 , and MlYAB2 were in the same subfamily (Figs. 1 B and S3 ). Significant elongation was also observed in the pistils (stigma) of OfYABBY12-OE tobacco flowers, while anther length remained unchanged.
The present study identified 13 OfYABBY genes in the O . fragrans genome, which were classified into five subfamilies. Phylogenetic analysis and gene duplication events demonstrated that gene duplication aided in the expansion of the O . fragrans OfYABBY gene family and that the gene functions of the OfYABBY family may be conserved. The genes in the YAB2, FIL/YAB3, and YAB5 subfamilies may have overlapping functions. In addition, the OfYABBY gene expression pattern analysis indicated that OfYABBY genes in the YAB2, FIL/YAB3, and YAB5 subfamilies may play important roles in O . fragrans leaf and/or floral organs. Functional validation showed that OfYABBY12 significantly affected the VOCs from tobacco leaves, especially β-ionone, and the dual-luciferase assay revealed that OfYABBY12 negatively regulated the expression of OfCCD4 , which promoted β-ionone synthesis. OfYABBY12 also played an important role in the establishment of polarity in tobacco leaves and in the development of lateral organs (pistils). The above results suggest that the OfYABBY gene family may participate in the growth and development of O . fragrans leaves and lateral organs, as well as in the synthesis of O . fragrans floral scent. In conclusion, this study provides a foundation for further research on the YABBY gene family in O . fragrans as well as new findings regarding the biosynthesis of the floral scent substance β-ionone in O . fragrans .
Data availability
All data generated or analyzed during this study are included in this published article, its supplementary information files and publicly available repositories. RNA-Seq raw data were uploaded to the NCBI sequence read archive ( http://www.ncbi.nlm.nih.gov/sra/ ) under accession number SRP450701 and are accessible under Bioproject archive number PRJNA997126 ( http://www.ncbi.nlm.nih.gov/bioproject/ ).
Wang LM, Li MT, Jin WW, Li S, Zhang SQ, Yu LJ. Variations in the components of Osmanthus fragrans Lour. Essential oil at different stages of flowering. Food Chem. 2009;114:233–6.
Article CAS Google Scholar
Zhou F, Zhao YJ, Li MQ, Xu T, Zhang LQ, Lu BY, Wu XD, Ge ZW. Degradation of phenylethanoid glycosides in Osmanthus fragrans Lour. Flowers and its effect on anti-hypoxia activity. Sci Rep. 2017;7:10068.
Article PubMed PubMed Central Google Scholar
Han Y, Dong MF, Yuan WJ, Shang FD. Study on the genetic diversity of Osmanthus fragrans cultivars. Chin Bull Bot. 2008;25:559–64.
CAS Google Scholar
Chen HG, Zeng XL, Yang J, Cai X, Shi YM, Zheng RR, Wang ZQ, Liu JY, Yi XX, Xiao SW, Fu Q, Zou JJ, Wang CY. Whole-genome resequencing of Osmanthus fragrans provides insights into flower color evolution. Hortic Res. 2021;8:98.
Article CAS PubMed PubMed Central Google Scholar
Feng YY, Li QY, Huang JH, Hu SQ. Numerical classification of 25 color-leafed Osmanthus fragrans clones (cultivars). J Nanjing Forestry Univ (Natural Sci Edition). 2021;45:107–15.
Google Scholar
Stahle MI, Kuehlich J, Staron L, von Arnim AG, Golz JF. YABBYs and the transcriptional corepressors LEUNIG and LEUNIG_HOMOLOG maintain leaf polarity and meristem activity in Arabidopsis. Plant Cell. 2009;21:3105–18.
Sarojam R, Sappl PG, Goldshmidt A, Efroni I, Floyd SK, Eshed Y, Bowman JL. Differentiating Arabidopsis shoots from leaves by combined YABBY activities. Plant Cell. 2010;22:2113–30.
Wang Q, Reddy VA, Panicker D, Mao HZ, Kumar N, Rajan C, Venkatesh PN, Chua NH, Sarojam R. Metabolic engineering of terpene biosynthesis in plants using a trichome-specific transcription factor MsYABBY5 from spearmint ( Mentha spicata ). Plant Biotechnol J. 2016;14:1619–32.
Zhang T, Li C, Li D, Liu Y, Yang X. Roles of YABBY transcription factors in the modulation of morphogenesis, development, and phytohormone and stress responses in plants. J Plant Res. 2020;133:751–63.
Article PubMed Google Scholar
Chen YY, Hsiao YY, Chang SB, Zhang D, Lan SR, Liu ZJ, Tsai WC. Genome-wide identification of YABBY genes in Orchidaceae and their expression patterns in Phalaenopsis orchid. Genes. 2020;11:955.
Eckardt NA. YABBY genes and the development and origin of seed plant leaves. Plant Cell. 2010; 22: 2103.
Satterlee JW, Scanlon MJ. Coordination of leaf development across developmental axes. Plants. 2019;8:433.
Zhang XL, Zhang LG. Molecular cloning and expression of the male sterility-related CtYABBY1 gene in flowering Chinese cabbage ( Brassica campestris L. Ssp chinensis var. Parachinensis ). Genet Mol Res. 2014;13:4336–47.
Article CAS PubMed Google Scholar
Strable J, Vollbrecht E. Maize YABBY genes drooping leaf1 and drooping leaf2 regulate floret development and floral meristem determinacy. Development. 2019;146:dev171181.
Zhang S, Wang L, Sun X, Li Y, Yao J, Nocker SV, Wang X. Genome-wide analysis of the YABBY gene family in grapevine and functional characterization of VvYABBY4 . Front Plant Sci. 2019;10:1207.
Jie GUO, Zhou XT, Dai KL, Yuan XY, Guo PY, Shi WP, Zhou MX. Comprehensive analysis of YABBY gene family in foxtail millet ( Setaria italica ) and functional characterization of SiDL . J Integr Agr. 2022;21:2876–87.
Article Google Scholar
Kayani SI, Shen Q, Ma YN, Fu XQ, Xie LH, Zhong YJ, Tiantian C, Pan QF, Li L, Rahman US, Sun XF, Tang KX. The YABBY family transcription factor AaYABBY5 directly targets cytochrome P450 monooxygenase (CYP71AV1) and double-bond reductase 2 (DBR2) involved in artemisinin biosynthesis in Artemisia Annua . Front Plant Sci. 2019;10:1084.
Kayani SI, Shen Q, Rahman SU, Fu XQ, Li YP, Wang C, Hassani D, Tang KX. Transcriptional regulation of flavonoid biosynthesis in Artemisia annua by AaYABBY5. Hortic Res. 2021;8:257.
Zhao SP, Lu D, Yu TF, Ji YJ, Zheng WJ, Zhang SX, Chai SC, Chen ZY, Cui XY. Genome-wide analysis of the YABBY family in soybean and functional identification of GmYABBY10 involvement in high salt and drought stresses. Plant Physiol Bioch. 2017;119:132–46.
Yang ZE, Gong Q, Wang LL, Jin YY, Xi JP, Li Z, Qin WQ, Yang ZR, Lu LL, Chen QJ, Li FG. Genome-wide study of YABBY genes in upland cotton and their expression patterns under different stresses. Front Genet. 2018;9:33.
Li ZY, Li G, Cai MX, Priyadarshani SV, Aslam M, Zhou Q, Huang XY, Wang XM, Liu YQ, Qin Y. Genome-wide analysis of the YABBY transcription factor family in pineapple and functional identification of AcYABBY4 involvement in salt stress. Int J Mol Sci. 2019;20:5863.
Bowman JL, Smyth DR. CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development. 1999;126:2387–96.
Villanueva JM, Broadhvest J, Hauser BA, Meister RJ, Schneitz K, Gasser CS. INNER NO OUTER regulates abaxial-adaxial patterning in Arabidopsis ovules. Gene Dev. 1999;13:3160–9.
Sun L, Wei YQ, Wu KH, Yan JY, Xu JN, Wu YR, Li GX, Xu JM, Harberd NP, Ding ZJ, Zheng SJ. Restriction of iron loading into developing seeds by a YABBY transcription factor safeguards successful reproduction in Arabidopsis . Mol Plant. 2021;14:1624–39.
Filyushin MA, Slugin MA, Dzhos EA, Kochieva EZ, Shchennikova AV. Coexpression of YABBY1 and YABBY3 genes in lateral organs of tomato species ( Solanum , Section Lycopersicon). Dokl Biochem Biophys. 2018;478:50–4.
Bartley GE, Ishida BK. Ethylene-sensitive and insensitive regulation of transcription factor expression during in vitro tomato sepal ripening. J Exp Bot. 2007;58:2043–51.
Yang T, He Y, Niu S, Zhang Y. A YABBY gene CRABS CLAW a ( CRCa ) negatively regulates flower and fruit sizes in tomato. Plant Sci. 2022;320:111285.
Zhang X, Ding L, Song AP, Li S, Liu JY, Zhao WQ, Jia DW, Guan YX, Zhao KK, Chen SM, Jiang JF, Chen FD. DWARF AND ROBUST PLANT regulates plant height via modulating gibberellin biosynthesis in chrysanthemum. Plant Physiol. 2022;190:2484–500.
Bonaccorso O, Lee JE, Puah L, Scutt CP, Golz JF. FILAMENTOUS FLOWER controls lateral organ development by acting as both an activator and a repressor. BMC Plant Biol. 2012;12:1–16.
Yang XL, Yue YZ, Li HY, Ding WJ, Chen GW, Shi TT, Chen JH, Park MS, Chen F, Wang LG. The chromosome-level quality genome provides insights into the evolution of the biosynthesis genes for aroma compounds of Osmanthus fragrans. Hortic Res. 2018;5:72.
Finn RD, Coggill PY, Eberhardt RY, Eddy SR, Mistry J, Mitchell AL, Potter SC, Punta M, Qureshi M, Sangrador-Vegas A, Salazar GA, Tate J, Bateman A. The pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 2016;44:D279–85.
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020;13:1194–202.
Shen Y, Li XM, Ma GL, Zhao Y, Jiang XL, Gao LP, Xia T, Liu YJ. Roles of YABBY tanscription factors in the regulation of leaf development and abiotic stress responses in Camellia sinensis . Bev Plant Res. 2022;2:1–10.
Ding BQ, Li JJ, Gurung VD, Lin QS, Sun XM, Yuan YW. The leaf polarity factors SGS3 and YABBYs regulate style elongation through auxin signaling in Mimulus lewisii . New Phytol. 2021;232:2191–206.
Li ZN, Jiang YJ, Liu DF, Ma J, Li J, Li MY, Sui SZ. Floral scent emission from nectaries in the adaxial side of the innermost and middle petals in Chimonanthus praecox . Int J Mol Sci. 2018;19:3278.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35:1547–9.
Zhang C, Fu JX, Wang YG, Bao ZY, Zhao HB. Identification of suitable reference genes for gene expression normalization in the quantitative real-time PCR analysis of sweet osmanthus (Osmanthus fragrans Lour). PLoS ONE. 2015;10:e0136355.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2. Method Methods. 2001;25:402–8.
Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews GN, Bowman JL. Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development. 1999;126:4117–28.
Yue YZ, Shi TT, Liu JW, Tian QY, Yang XL, Wang LG. Genomic, metabonomic and transcriptomic analyses of sweet osmanthus varieties provide insights into floral aroma formation. Sci Hortic. 2022;306:111442.
Huang FC, Molnar P, Schwab W. Cloning and functional characterization of carotenoid cleavage dioxygenase 4 genes. J Exp Bot. 2009;60:3011–22.
Zhang XS, Pei JS, Zhao LG, Tang F, Fang XY, Xie JC. Overexpression and characterization of CCD4 from Osmanthus fragrans and β-ionone biosynthesis from β-carotene in vitro. J Mol Catal B-Enzym. 2016;134:105–14.
Han HQ, Liu Y, Jiang MM, Ge HY, Chen HY. Identification and expression analysis of YABBY family genes associated with fruit shape in tomato ( Solanum lycopersicum L). Genet Mol Res. 2015;14:7079–91.
Bowman JL. The YABBY gene family and abaxial cell fate. Curr Opin Plant Biol. 2000;3:17–22.
Wang AJ, Tang JF, Li DY, Chen CY, Zhao XY, Zhu LH. Isolation and functional analysis of LiYAB1 , a YABBY family gene, from lily ( Lilium longiflorum ). J Plant Physiol. 2009;166:988–95.
Du F, Guan C, Jiao Y. Molecular mechanisms of leaf morphogenesis. Mol Plant. 2018;11:1117–34.
Wang HF, Kong FJ, Zhou CE. From genes to networks: the genetic control of leaf development. J Integr Plant Biol. 2021;63:1181–96.
Boter M, Golz JF, Giménez-Ibañez S, Fernandez-Barbero G, Franco-Zorrilla JM, Solano R. FILAMENTOUS FLOWER is a direct target of JAZ3 and modulates responses to jasmonate. Plant Cell. 2015;27:3160–74.
Kumaran MK, Bowman JL, Sundaresan V. YABBY polarity genes mediate the repression of KNOX homeobox genes in Arabidopsis. Plant Cell. 2002;14:2761–70.
Garrido-Bigotes A, Torrejón M, Solano R, Figueroa CR. Interactions of JAZ repressors with anthocyanin biosynthesis-related transcription factors of Fragaria × ananassa . Agronomy. 2020;10:1586.
Goldshmidt A, Alvarez JP, Bowman JL, Eshed Y. Signals derived from YABBY gene activities in organ primordia regulate growth and partitioning of Arabidopsis shoot apical meristems. Plant Cell. 2008;20:1217–30.
Han YJ, Wang HY, Wang XD, Li K, Dong MF, Li Y, Zhu Q, Shang FD. Mechanism of floral scent production in Osmanthus fragrans and the production and regulation of its key floral constituents, β-ionone and linalool. Hortic Res. 2019;6:106.
She ZY, Huang XY, Aslam M, Wang LL, Yan MK, Qin RJ, Chen YZ, Qin Y, Niu XP. Expression characterization and cross-species complementation uncover the functional conservation of YABBY genes for leaf abaxial polarity and carpel polarity establishment in Saccharum spontaneum . BMC Plant Biol. 2022;22:124.
Download references
Acknowledgements
We would like to thank all of the colleagues in our laboratory for providing useful discussions and technical assistance.
This work was supported by the National Natural Science Foundation of China (32071828) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and affiliations.
College of Landscape Architecture, Nanjing Forestry University, Nanjing, Jiangsu Province, 210037, China
Tingting Shi, Ling Zhou, Yunfang Ye, Xiulian Yang, Lianggui Wang & Yuanzheng Yue
Co-Innovation Center for Sustainable Forestry in Southern China, State Key Laboratory of Tree Genetics and Breeding, Nanjing Forestry University, Nanjing, 210037, China
You can also search for this author in PubMed Google Scholar
Contributions
T.T.S. and Y.Z.Y. conceived and designed the research; T.T.S. and Y.F.Y. performed all experiments; X.L.Y. and L.G.W. provided technical assistance; T.T.S. wrote the manuscript; and L. Z., Y.Z.Y., and L.G.W. revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Correspondence to Lianggui Wang or Yuanzheng Yue .
Ethics declarations
Ethics approval and consent to participate.
Not applicable.
Consent for publication
Competing interests.
The authors declare no competing interests.
Additional information
Publisher’s note.
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1
Supplementary material 2, rights and permissions.
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/ . The Creative Commons Public Domain Dedication waiver ( http://creativecommons.org/publicdomain/zero/1.0/ ) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
Reprints and permissions
About this article
Cite this article.
Shi, T., Zhou, L., Ye, Y. et al. Characterization of YABBY transcription factors in Osmanthus fragrans and functional analysis of OfYABBY12 in floral scent formation and leaf morphology. BMC Plant Biol 24 , 589 (2024). https://doi.org/10.1186/s12870-024-05047-y
Download citation
Received : 16 November 2023
Accepted : 19 April 2024
Published : 21 June 2024
DOI : https://doi.org/10.1186/s12870-024-05047-y

Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Osmanthus fragrans
- YABBY gene family
- Leaf development
- Volatile organic compounds
BMC Plant Biology
ISSN: 1471-2229
- General enquiries: [email protected]
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- Front Immunol
- PMC10034027
Transcription factors in megakaryocytes and platelets
Hengjie yuan.
1 Tianjin Institute of Neurology, Department of Neurosurgery, Tianjin Medical University General Hospital, Tianjin, China
2 BloodWorks Research Institute, Seattle, WA, United States
Jianning Zhang
Jing-fei dong.
3 Division of Hematology, Department of Medicine, University of Washington, School of Medicine, Seattle, WA, United States
Zilong Zhao
Transcription factors bind promoter or regulatory sequences of a gene to regulate its rate of transcription. However, they are also detected in anucleated platelets. The transcription factors RUNX1, GATA1, STAT3, NFκB, and PPAR have been widely reported to play key roles in the pathophysiology of platelet hyper-reactivity, thrombosis, and atherosclerosis. These non-transcriptional activities are independent of gene transcription or protein synthesis but their underlying mechanisms of action remain poorly defined. Genetic and acquired defects in these transcription factors are associated with the production of platelet microvesicles that are known to initiate and propagate coagulation and to promote thrombosis. In this review, we summarize recent developments in the study of transcription factors in platelet generation, reactivity, and production of microvesicles, with a focus on non-transcriptional activities of selected transcription factors.
1. Introduction
Transcription factors (TFs) are a group of mediators that bind the promoter or regulatory sequence of a gene to control its rate of transcribing genetic information from DNA to messenger RNA ( 1 ). This transcription control is key to ensuring an adequate level of expression of a given protein in targeted cells at a particular developmental stage. It not only directs the processes of proliferation, growth, and death of a cell, but also controls the rate of cell migration and organizational development during embryonic development, as well as regulating cellular response to the extracellular matrices. Thus far, more than 1600 transcription factors have so far been identified ( 2 , 3 ), and they work in a coordinated fashion to down- as well as up-regulate target genes. The activation of a given gene can be regulated by multiple transcriptional factors and one transcription factor can regulate multiple genes. Such a multivalent activity is possible because of the modular structure of a transcriptional factor, which typically includes a DNA-binding domain, signal-sensing domain that contains binding sites for transcription co-regulators, and an optional transactivation domain, which senses external signals and transmits them to the rest of the transcription complex ( 4 , 5 ). Because of their roles in regulating gene transcription, the activation and suppression of transcription factors is extensively reported in cancer development ( 6 ).
Paradoxically, multiple transcription factors have been reported to express and be active in platelets ( Table 1 ), the anucleated offspring of megakaryocytes with a very limited capacity for protein synthesis ( 7 ). An obvious question is whether these transcription factors are merely leftover from parental megakaryocytes or have unique activities in platelets. Reports from studies on platelet transcription factors have been scarce in the literature, but increasing evidence suggests that transcription factors in platelets have unique activities of their own independent of their transcriptional activities ( 8 – 10 ). However, past research on transcription factors in platelets is often limited to reporting their presence and activation status, without further investigation of their activities in regulating platelet functions and, more importantly the underlying mechanism of their regulatory activities.
Table 1
Roles of transcription factors in platelets.
| Transcription factor | Roles under activation or mutation | Associated hematologic abnormalities |
|---|---|---|
| RUNX1 | Platelet granule development, platelet activation | MDS/AML |
| GATA1 | Inhibit aggregation | Dyserythropoiesis |
| STAT3 | Increase aggregation, P-selectin, thrombosis | Coronary artery diseases |
| NFkB | Increase aggregation, spreading, clot retraction, GPIBa shedding | Cardiovascular diseases |
| PPAR | Inhibition of platelet function | Cardiovascular diseases |
AML, acute myeloid leukemia; MDS, myelodysplastic syndrome.
Platelets circulate along the vessel wall and act to stop bleeding at sites of vessel injury. This hemostatic process requires multiple ligand-receptor interactions to tether, activate, and aggregate platelets. The tightly controlled platelet activation and aggregation that occurs at the site of vascular injury during hemostasis can become dysregulated in pathological conditions, promoting thrombosis and inflammation. For example, platelets promote arterial thrombosis or thromboembolism when activated either on the surface of a ruptured atherosclerotic plaque or by pathological levels of high fluid shear stress in the area of arterial stenosis, leading to acute thrombotic events such as ischemic stroke and myocardial infarction ( 11 ). Emerging evidence further suggests that platelets also act as a cellular mediator in a variety of pathophysiological conditions such as cancer, rheumatoid arthritis, atherosclerosis, trauma, and immune response ( 12 – 14 ). How transcription factors regulate platelet production from megakaryocytes has been extensively reported, but their non-transcriptional activities (i.e., activity independent of gene regulations) have only begun to be recognized. Here, we discuss several transcription factors that have been reported to regulate platelet production and function.
2. Transcription factors in platelet production
2.1. runt-related transcription factor 1.
In 1969, Weiss, et al. identified a family with an autosomal dominant inherited thrombocytopenia, caused primarily by decreased dense granule contents ( 15 ). A heterozygous Y260X mutation in the RUNX1 gene was subsequently shown to be the genetic basis of this inherited platelet defect ( 15 , 16 ). To date, more than 200 families with RUNX1 variants have been reported ( 17 ). RUNX1/AML1 (also known as CBFA2 and PEBP2αB) is a member of the Runt family, which has three known transcription factors (RUNX1, RUNX2, and RUNX3), which share the Runt homology domain near the N-terminus. This domain interacts with CBFb to bind specific sequences of DNA to regulate its transcription ( 18 ).
RUNX1 regulates several genes that control platelet production, structure, function, and intracellular signaling. One report found that 22 patients in a family with autosomal dominant thrombocytopenia had mutations in the RUNX1 gene ( 19 ) and 6 of them developed hematologic malignancies ( 20 ). RUNX1-deficient mice die in uterus due to defective hematopoiesis and resultant severe bleeding ( 21 , 22 ). Mice with the conditional knockout survive but have an impaired megakaryocyte maturation with a significant reduction in megakaryocyte polyploidization ( 23 ). Variations in the RUNX1 gene often result in bleeding diathesis, primarily because of defective platelet granules ( 15 , 16 ), which reduce platelet activation and aggregation ( 24 ). For example, mice carrying the RUNX1 p.Leu43Ser variant (equivalent to human p.Leu56Ser) exhibit a prolonged bleeding time because of defective α-granule secretion and platelet spreading ( 25 ). RUNX1 deficiency can result in pallidin dysregulation and deficient dense granules in platelets ( 26 ) as well as the Ras-related protein RAB31-mediated early endosomal trafficking of von Willebrand factor (VWF) and epidermal growth factor receptor (EGFR) in megakaryocytes ( 27 ). RUNX1 regulates the development of platelet granules through interaction with genes involved in the biogenesis of platelet granules such as the nuclear factor erythroid 2 (NF-E2).
In addition, RUNX1 can also regulate genes related to platelet functions. For example, it regulates the transcription of the non-muscle myosin IIA (MYH9) and IIB (MYH10) genes, which encode non-muscle myosin II heavy chains; RUNX1 mutations are associated with dysregulated expression of MYH10 in platelets ( 28 ); and the expression level of non-muscle myosin is used as a marker for changes in transcriptional activity of RUNX1 as well as friend leukemia integration 1 transcription factor (FLI1) ( 29 ). RUNX1 also regulates the expression of the arachidonate 12-lipoxygenase gene (ALOX12) ( 30 ), which encodes the enzyme that acts on polyunsaturated fatty acid substrates to generate bioactive lipid mediators to regulate platelet function ( 30 ). PCTP (phosphatidylcholine transfer protein) regulates the intermembrane transfer of phosphatidylcholine and its upregulation by RUNX1 sensitizes platelet response to thrombin through protease-activated receptor 4 ( 31 ). RUNX1 also regulates the expression of platelet factor 4 through coordination with transcription factors in the ETS family that share a conserved winged helix-turn-helix DNA binding domain that recognizes unique DNA sequences containing GGAA/T ( 32 ). Platelet factor 4 belongs to the CXC chemokine family and is released from α-granules of activated platelets to promote coagulation and to participate in heparin-induced thrombocytopenia ( 33 , 34 ). A recent report shows that RUNX-1 haploinsufficiency inhibits the differentiation of hematopoietic progenitor cells (HPCs) into megakaryocytes ( 35 ).
2.2. GATA-binding protein 1
GATA-binding protein 1 (GATA1) is a transcription factor that contains two zinc finger domains: a C-terminal zinc finger that binds the (T/A) GATA(A/G) motif of DNA and an N-terminal zinc finger that is required for stabilizing the C-terminal structure and also interacts with a nuclear co-factor protein called friend for GATA1 (FOG1), which stabilizes GATA1 binding ( 36 , 37 ). GATA plays a pivotal role in hematopoietic development and is found in megakaryocytes ( 38 ). GATA1-deficient mice die before birth at approximately embryonic day 10, primarily because of severe anemia ( 39 ). However, mutations in the N-terminal zinc finger domain, which reduces the transcriptional activation of GATA1 ( 36 , 40 ), are found in patients with myeloproliferative disorders and acute megakaryoblastic leukemia ( 41 ), suggesting that GATA1-FOG1 interaction is essential for the development and maturation of megakaryocytes, the parental cells of platelets. Decreased GATA-1 expression has also been reported in patients with myelodysplastic syndrome ( 42 ).
Embryonic stem cells from GATA1-deficient mice are smaller and show low expression of megakaryocytic markers, but have a high rate of proliferation ( 43 ). Complementation of these cells with a wild-type GATA1 gene allows megakaryocytes and erythrocytes to develop in response to a variety of cytokines. Additionally, cell division is attenuated in the megakaryocytic progenitor G1ME cells that overexpress GATA1. A recent report further shows that impaired MYH10 silencing causes GATA1-related polyploidization defect during megakaryocyte differentiation ( 44 ).
Furthermore, platelet aggregation induced by collagen is inhibited in GATA1 - deficient mice ( 45 ), primarily due to reduced expression of the collagen receptor GPVI. Platelet adhesion and aggregation induced by shear stress are also reduced in GATA1 - deficient mice ( 45 ). How a GATA1 deficiency causes these changes in platelet reactivity remains unknown, but these phenotypic changes in the mice provide the first indication that transcription factors could perform non-transcriptional activities in anucleated platelets.
3. Non-transcriptional activity in platelets
3.1. signal transducer and activator of transcription 3.
STAT includes a family of transcription factors critical for inflammatory and acute-phase reactions ( 46 , 47 ). They also play vital roles in cancer development and hematopoiesis ( 48 ). The homologous STAT1, STAT3, and STAT5 are expressed in human platelets and are reported to regulate platelet reactivity through residual or mitochondrial transcriptional activity in platelets. For example, STAT3 affects mitochondrial transcription by binding to the regulatory D-loop region of mitochondrial DNA upon platelet activation ( 49 ).
However, STAT3 can also be activated (phosphorylated) and dimerized in platelets stimulated with thrombopoietin ( 49 , 50 ), suggesting that STAT3 can also regulate platelet reactivity through non-transcriptional means. We have shown that STAT3 is activated and dimerized in collagen-stimulated platelets to serve as a protein scaffold that facilitates the catalytic interaction between spleen tyrosine kinase (Syk) and its substrate, PLCγ to enhance collagen-induced calcium mobilization and platelet activation ( 8 ). More importantly, STAT3 is activated to form dimers by a complex of IL-6 with its soluble receptor IL-6Rα, which activates JAK2 ( 51 ). The pharmacological inhibition of platelet STAT3 reduces collagen-induced platelet aggregation and thrombus formation on the collagen matrix ( 8 , 52 ). Platelets from STAT3-deficient mice or mice infused with a STAT3 inhibitor have reduced collagen-induced aggregation. This non-transcriptional activity of STAT3 may be critical for the development of platelet hyper-reactivity, which has been widely associated with inflammation, especially that related to the activity of the proinflammatory cytokine IL-6 ( 8 ). We have also shown that the piper longum derivative piperlongumine (PL) blocks collagen-induced platelet reactivity in a dose-dependent manner by targeting STAT3 ( 53 ). Consistent with our observations, the small molecular STAT3 inhibitor SC99 has been shown to reduce platelet activation and aggregation induced by collagen and thrombin ( 54 ). These findings offer a new pathway for reducing platelet hyper-reactivity in conditions of inflammation and in prothrombotic states associated with trauma, cancer, autoimmune diseases, and severe infection.
3.2. Nuclear factor kappa β
Nuclear factor kappa β (NFκB) is a well-defined redox-sensitive transcription factor that regulates the immune response and inflammation by controlling the expression of multiple genes activated by inflammatory mediators ( 55 – 57 ). Blocking NFκB can therefore improve outcomes of inflammatory diseases ( 58 ). NFκB is composed of p50 and p65 subunits, normally as an inactive cytoplasmic complex. The inhibitory proteins of the IκB family tightly bind the subunits of NFκB ( 59 ). Upon activation, the IκK complex phosphorylates IκBα, thus activating NFκB by detaching it from IkBα ( 60 – 62 ). Three IκK family members, α, β, and γ, are expressed in platelets, with β being the most abundant, and are reported to regulate platelet reactivity through non-transcriptional activity ( 9 , 10 , 63 ). For example, the pharmacological inhibition of IκKβ leads to reduced agonist-induced platelet activation, increased bleeding time, and prolonged thrombus formation in a mouse model ( 64 ). NF-κB has also been reported to be partially involved in the regulation of SERCA activity to regulate calcium homeostasis in platelets ( 65 ). IκKβ-deficient platelets lose the ability to shed the ectodomain of GP Ibα in response to ADP or collagen stimulations ( 66 ) but preserve thrombin-induced GP Ibα shedding ( 67 ). Collagen-induced p65 and IκKβ phosphorylation is blocked by inhibition of MAP kinase, but not by inhibition of ERK in platelets ( 68 ). The thrombin-induced GP Ibα shedding requires p38 mitogen-activated protein kinase (MAPK) and extracellular signal-regulated kinase (ERK) as its upstream and downstream molecules ( 68 , 69 ).
3.3. Peroxisome proliferator-activated receptors
The peroxisome proliferator-activated receptors (PPARs) are ligand-activated receptors in the nuclear hormone receptor family. They contain three subtypes (PPARα, PPARβ/δ, and PPARγ), which are essential in the regulation of cell differentiation, development, and metabolism ( 70 – 72 ). All PPARs heterodimerize with retinoid X receptor (RXR) and subsequently bind to a specific region of target genes called a peroxisome proliferator response element (PPRE) ( 73 ). PPARγ plays a transcription factor role in regulating platelet production from megakaryocytes, but the PPARγ ligand thiazolidinedione inhibits platelet aggregation induced by ADP under hydrostatic pressure and in diabetic mice ( 74 – 76 ). Similarly, activating PPARβ/δ also reduces platelet reactivity to ADP, thrombin, and collagen ( 77 , 78 ). However, PPARα is also required for platelet activation and thrombus formation, in which it regulates the dense granule secretion of platelets in hyperlipidemic mice ( 79 ). The reason for this apparent contradiction remains to be further investigated. PPARγ is recruited and phosphorylated by Syk to promote the recruitment of the protein called Linker for the Activation of T cells (LAT), which is necessary for collagen-induced platelet activation through glycoprotein VI ( 80 ).
While transcription factors are critically involved in megakaryocyte development and platelet production, they may also regulate platelet reactivity to conventional and specific platelet agonists ( Figure 1 ). The latter is independent of transcriptional activity, for which it is present but at a residual level. This non-transcriptional activity remains poorly understood and requires further investigation because it helps understanding how platelets are activated either by conventional agonists for hemostasis or as complications found in patients treated with drugs that block transcriptional activity of cells (e.g., cancer treatments). Such research will also play an important role in developing new therapeutics targeting these transcription factors to enhance or reduce platelet reactivity.

Transcription factors regulate platelet aggregation through non-transcriptional activities. (A) PPARγ is recruited and phosphorylated by Syk to promote the recruitment of LAT and enhance platelet aggregation; (B) NFκB is activated by upstream p38 mitogen-activated protein kinase (MAPK) and promotes platelet aggregation by regulating downstream extracellular signal-regulated kinase (ERK); (C) A complex of IL-6 with its soluble receptor IL-6R activates JAK2 to phosphorylate and dimerize STAT3, then the activated STAT3 serves as a protein scaffold to facilitate the catalytic interaction between the spleen tyrosine kinase (Syk) and its substrate PLCγ2 to promote platelet aggregation.
4. Transcription factors in extracellular vesicles released from platelets
Extracellular vesicles (EVs) are shed membrane fragments, intracellular organelles, and nuclear components from cells undergoing active microvesiculation ( 81 – 84 ) or apoptosis ( 85 – 87 ). The former is triggered by the activation of the cysteine protease calpain, which disrupts the membrane-cytoskeleton association ( 88 – 91 ). Platelets are the primary source of EVs circulating in blood, accounting for approximately 80% of total EVs ( 92 – 94 ). The subcellular size of EVs allows them to travel to areas where parental cells are unable to go. In additional to inherent functions from their parental cells, EVs also perform unique activities of their own because of molecules expressed on their surface and carried by them, the latter of which include transcription factors such as STAT3, STAT5, and PPARγ ( 95 ) as well as regulators of transcription factors ( 96 , 97 ). This EV-derived transcriptional activity has been scarcely reported but hold greats potential for influencing biological activities of target cells. For example, PPARγ in platelet EVs is taken up by monocytic THP-1 cells, where it induces the expression of fatty acid-binding protein-4 (FABP4). Monocytes receiving PPARγ-containing platelet EVs produce less inflammatory mediators and become more adherent through increased fibronectin production ( 95 ). Although reports on platelet-derived transcription factors remain very limited, a large body of evidence in the literature shows that platelet-derived EVs, especially EV-carried microRNAs, can change transcriptional activities, thus regulating the function of target cells. Platelet EV-carried NLR family pyrin domain containing 3 (NLRP3) stimulates endothelial cells to undergo pyroptosis through the NLRP3/nuclear factor (NF)-κB pathway ( 98 ). EVs from platelets stimulated with bacteria provoke proinflammatory activity of monocytes through the TRAF6/NFκB pathway ( 99 ). MicroRNA-142-3p carried by platelet-derived EVs promotes the proliferation of endothelial cells ( 100 ), whereas microRNA-126-3p-carrying platelet EVs can be internalized by macrophages to dose-dependently downregulate expression of target mRNA ( 101 ). These observations mostly pertain to phenotypic characterization with less information regarding the underlying pathways involved. Systemic studies of EV-carrying transcription factors and related mediators are therefore urgently needed.
5. Conclusion
Platelets lack a nucleus and de novo transcription, but a number of transcription factors are found in platelets and may have non-transcriptional activities that regulate platelet function. Transferring transcription factors between platelets and target cells through platelet EVs could also be a novel regulatory mechanism of cell-cell communications and a potential therapeutic target for a variety of pathologies.
Author contributions
HY and YL performed the literature search and compiled all the information from the researched articles and wrote the manuscript. ZZ, J-FD and JZ formulated, proposed, guided and wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding Statement
This study is supported by Young Scientists Award 82022020 from the National Natural Science Foundation of China (ZZ), National Natural Science Foundation of China 81971176 (ZZ), 81271361, 81271359 (JZ), 81102447 (HY), National Natural Science Foundation of China State Key Program Grant 81330029, National Natural Science Foundation of China Major International Joint Research Project 81720108015 (JZ), and Postdoctoral Science Foundation of China Grants 2013M541190 (HY).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Advertisement

Single-Cell Chromatin Accessibility Analysis Reveals the Epigenetic Basis and Signature Transcription Factors for the Molecular Subtypes of Colorectal Cancers
Z. Liu, Y. Hu, H. Xie, and K. Chen contributed equally to this article.
Cancer Discov 2024;14:1082–105
- Funder(s): National Natural Science Foundation of China (NSFC)
- Award Id(s): 81972702
- Funder(s): Beijing Nova Program
- Award Id(s): 2022029
- Article contents
- Figures & tables
- Supplementary Data
- Peer Review
- Get Permissions
- Cite Icon Cite
- Search Site
- Version of Record June 3 2024
- Proof April 4 2024
- Accepted Manuscript March 6 2024
Zhenyu Liu , Yuqiong Hu , Haoling Xie , Kexuan Chen , Lu Wen , Wei Fu , Xin Zhou , Fuchou Tang; Single-Cell Chromatin Accessibility Analysis Reveals the Epigenetic Basis and Signature Transcription Factors for the Molecular Subtypes of Colorectal Cancers. Cancer Discov 1 June 2024; 14 (6): 1082–1105. https://doi.org/10.1158/2159-8290.CD-23-1445
Download citation file:
- Ris (Zotero)
- Reference Manager
Colorectal cancer is a highly heterogeneous disease, with well-characterized subtypes based on genome, DNA methylome, and transcriptome signatures. To chart the epigenetic landscape of colorectal cancers, we generated a high-quality single-cell chromatin accessibility atlas of epithelial cells for 29 patients. Abnormal chromatin states acquired in adenomas were largely retained in colorectal cancers, which were tightly accompanied by opposite changes of DNA methylation. Unsupervised analysis on malignant cells revealed two epigenetic subtypes, exactly matching the iCMS classification, and key iCMS-specific transcription factors (TFs) were identified, including HNF4A and PPARA for iCMS2 tumors and FOXA3 and MAFK for iCMS3 tumors. Notably, subtype-specific TFs bind to distinct target gene sets and contribute to both interpatient similarities and diversities for both chromatin accessibilities and RNA expressions. Moreover, we identified CpG-island methylator phenotypes and pinpointed chromatin state signatures and TF regulators for the CIMP-high subtype. Our work systematically revealed the epigenetic basis of the well-known iCMS and CIMP classifications of colorectal cancers.
Our work revealed the epigenetic basis of the well-known iCMS and CIMP classifications of colorectal cancers. Moreover, interpatient minor similarities and major diversities of chromatin accessibility signatures of TF target genes can faithfully explain the corresponding interpatient minor similarities and major diversities of RNA expression signatures of colorectal cancers, respectively.
This article is featured in Selected Articles from This Issue, p. 897
Client Account
Citing articles via, email alerts.
- Online First
- Online ISSN 2159-8290
- Print ISSN 2159-8274
AACR Journals
- Blood Cancer Discovery
- Cancer Discovery
- Cancer Epidemiology, Biomarkers & Prevention
- Cancer Immunology Research
- Cancer Prevention Research
- Cancer Research
- Cancer Research Communications
- Clinical Cancer Research
- Molecular Cancer Research
- Molecular Cancer Therapeutics
- Info for Advertisers
- Information for Institutions/Librarians
- Privacy Policy
- Copyright © 2023 by the American Association for Cancer Research.
This Feature Is Available To Subscribers Only
Sign In or Create an Account
- Search Menu
Sign in through your institution
- Advance Articles
- Collections
- Focus Collections
- Browse by cover
- High-Impact Research
- Author Guidelines
- Quick and Simple Author Support
- Focus Issues Call for Papers
- Submission Site
- Open Access Options
- Self-Archiving Policy
- Why Publish with Us?
- About Plant Physiology
- Editorial Board
- Advertising & Corporate Services
- Journals on Oxford Academic
- Books on Oxford Academic

- < Previous
Transcription factor CaHDZ15 promotes pepper basal thermotolerance by activating HEAT SHOCK FACTORA6a
- Article contents
- Figures & tables
- Supplementary Data
Shaoliang Mou, Weihong He, Haitao Jiang, Qianqian Meng, Tingting Zhang, Zhiqin Liu, Ailian Qiu, Shuilin He, Transcription factor CaHDZ15 promotes pepper basal thermotolerance by activating HEAT SHOCK FACTORA6a , Plant Physiology , Volume 195, Issue 1, May 2024, Pages 812–831, https://doi.org/10.1093/plphys/kiae037
- Permissions Icon Permissions
High temperature stress (HTS) is a serious threat to plant growth and development and to crop production in the context of global warming, and plant response to HTS is largely regulated at the transcriptional level by the actions of various transcription factors (TFs). However, whether and how homeodomain-leucine zipper (HD-Zip) TFs are involved in thermotolerance are unclear. Herein, we functionally characterized a pepper ( Capsicum annuum ) HD-Zip I TF CaHDZ15. CaHDZ15 expression was upregulated by HTS and abscisic acid in basal thermotolerance via loss- and gain-of-function assays by virus-induced gene silencing in pepper and overexpression in Nicotiana benthamiana plants. CaHDZ15 acted positively in pepper basal thermotolerance by directly targeting and activating HEAT SHOCK FACTORA6a ( HSFA6a ), which further activated CaHSFA2 . In addition, CaHDZ15 interacted with HEAT SHOCK PROTEIN 70-2 (CaHsp70-2) and glyceraldehyde-3-phosphate dehydrogenase1 (CaGAPC1), both of which positively affected pepper thermotolerance. CaHsp70-2 and CaGAPC1 promoted CaHDZ15 binding to the promoter of CaHSFA6a , thus enhancing its transcription. Furthermore, CaHDZ15 and CaGAPC1 were protected from 26S proteasome-mediated degradation by CaHsp70-2 via physical interaction. These results collectively indicate that CaHDZ15, modulated by the interacting partners CaGAPC1 and CaHsp70-2, promotes basal thermotolerance by directly activating the transcript of CaHSFA6a . Thus, a molecular linkage is established among CaHsp70-2, CaGAPC1, and CaHDZ15 to transcriptionally modulate CaHSFA6a in pepper thermotolerance.

American Society of Plant Biologists members
Personal account.
- Sign in with email/username & password
- Get email alerts
- Save searches
- Purchase content
- Activate your purchase/trial code
- Add your ORCID iD
Institutional access
Sign in with a library card.
- Sign in with username/password
- Recommend to your librarian
- Institutional account management
- Get help with access
Access to content on Oxford Academic is often provided through institutional subscriptions and purchases. If you are a member of an institution with an active account, you may be able to access content in one of the following ways:
IP based access
Typically, access is provided across an institutional network to a range of IP addresses. This authentication occurs automatically, and it is not possible to sign out of an IP authenticated account.
Choose this option to get remote access when outside your institution. Shibboleth/Open Athens technology is used to provide single sign-on between your institution’s website and Oxford Academic.
- Click Sign in through your institution.
- Select your institution from the list provided, which will take you to your institution's website to sign in.
- When on the institution site, please use the credentials provided by your institution. Do not use an Oxford Academic personal account.
- Following successful sign in, you will be returned to Oxford Academic.
If your institution is not listed or you cannot sign in to your institution’s website, please contact your librarian or administrator.
Enter your library card number to sign in. If you cannot sign in, please contact your librarian.
Society Members
Society member access to a journal is achieved in one of the following ways:
Sign in through society site
Many societies offer single sign-on between the society website and Oxford Academic. If you see ‘Sign in through society site’ in the sign in pane within a journal:
- Click Sign in through society site.
- When on the society site, please use the credentials provided by that society. Do not use an Oxford Academic personal account.
If you do not have a society account or have forgotten your username or password, please contact your society.
Sign in using a personal account
Some societies use Oxford Academic personal accounts to provide access to their members. See below.
A personal account can be used to get email alerts, save searches, purchase content, and activate subscriptions.
Some societies use Oxford Academic personal accounts to provide access to their members.
Viewing your signed in accounts
Click the account icon in the top right to:
- View your signed in personal account and access account management features.
- View the institutional accounts that are providing access.
Signed in but can't access content
Oxford Academic is home to a wide variety of products. The institutional subscription may not cover the content that you are trying to access. If you believe you should have access to that content, please contact your librarian.
For librarians and administrators, your personal account also provides access to institutional account management. Here you will find options to view and activate subscriptions, manage institutional settings and access options, access usage statistics, and more.
Short-term Access
To purchase short-term access, please sign in to your personal account above.
Don't already have a personal account? Register
| Month: | Total Views: |
|---|---|
| January 2024 | 141 |
| February 2024 | 158 |
| March 2024 | 185 |
| April 2024 | 67 |
| May 2024 | 74 |
| June 2024 | 39 |
Email alerts
Citing articles via.
- Recommend to Your Librarian
- Advertising & Corporate Services
- Awards & Funding
- Plant Science Today
- Plant Biology Meeting
- Meeting Management Services
- Plant Science Research Weekly
- Taproot: A Plantae Podcast
Affiliations
- Online ISSN 1532-2548
- Print ISSN 0032-0889
- Copyright © 2024 American Society of Plant Biologists
- About Oxford Academic
- Publish journals with us
- University press partners
- What we publish
- New features
- Open access
- Rights and permissions
- Accessibility
- Advertising
- Media enquiries
- Oxford University Press
- Oxford Languages
- University of Oxford
Oxford University Press is a department of the University of Oxford. It furthers the University's objective of excellence in research, scholarship, and education by publishing worldwide
- Copyright © 2024 Oxford University Press
- Cookie settings
- Cookie policy
- Privacy policy
- Legal notice
This Feature Is Available To Subscribers Only
Sign In or Create an Account
This PDF is available to Subscribers Only
For full access to this pdf, sign in to an existing account, or purchase an annual subscription.
Information
- Author Services
Initiatives
You are accessing a machine-readable page. In order to be human-readable, please install an RSS reader.
All articles published by MDPI are made immediately available worldwide under an open access license. No special permission is required to reuse all or part of the article published by MDPI, including figures and tables. For articles published under an open access Creative Common CC BY license, any part of the article may be reused without permission provided that the original article is clearly cited. For more information, please refer to https://www.mdpi.com/openaccess .
Feature papers represent the most advanced research with significant potential for high impact in the field. A Feature Paper should be a substantial original Article that involves several techniques or approaches, provides an outlook for future research directions and describes possible research applications.
Feature papers are submitted upon individual invitation or recommendation by the scientific editors and must receive positive feedback from the reviewers.
Editor’s Choice articles are based on recommendations by the scientific editors of MDPI journals from around the world. Editors select a small number of articles recently published in the journal that they believe will be particularly interesting to readers, or important in the respective research area. The aim is to provide a snapshot of some of the most exciting work published in the various research areas of the journal.
Original Submission Date Received: .
- Active Journals
- Find a Journal
- Proceedings Series
- For Authors
- For Reviewers
- For Editors
- For Librarians
- For Publishers
- For Societies
- For Conference Organizers
- Open Access Policy
- Institutional Open Access Program
- Special Issues Guidelines
- Editorial Process
- Research and Publication Ethics
- Article Processing Charges
- Testimonials
- Preprints.org
- SciProfiles
- Encyclopedia

Article Menu

- Subscribe SciFeed
- Recommended Articles
- Google Scholar
- on Google Scholar
- Table of Contents
Find support for a specific problem in the support section of our website.
Please let us know what you think of our products and services.
Visit our dedicated information section to learn more about MDPI.
JSmol Viewer
Interaction of the transcription factors bes1/bzr1 in plant growth and stress response.

1. Introduction
2. interactors of bes1/bzr1 in plant growth and development, 2.1. interactors of bes1/bzr1 in skotomorphogenesis and photomorphogenesis, 2.2. interactors of bes1/bzr1 in root growth, 2.3. interactors of bes1/bzr1 in other developmental processes, 3. interactors of bes1/bzr1 in stress response, 3.1. interactors of bes1/bzr1 in abiotic stress response, 3.2. interactors of bes1/bzr1 in biotic stress response, 4. conclusions, author contributions, institutional review board statement, informed consent statement, data availability statement, conflicts of interest.
- Li, J.; Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 1997 , 90 , 929–938. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Wang, Z.Y.; Seto, H.; Fujioka, S.; Yoshida, S.; Chory, J. BRI1 is a critical component of a plasma-membrane receptor for plant steroids. Nature 2001 , 410 , 380–383. [ Google Scholar ] [ CrossRef ]
- Kinoshita, T.; Cano-Delgado, A.; Seto, H.; Hiranuma, S.; Fujioka, S.; Yoshida, S.; Chory, J. Binding of brassinosteroids to the extracellular domain of plant receptor kinase BRI1. Nature 2005 , 433 , 167–171. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Cano-Delgado, A.; Yin, Y.; Yu, C.; Vafeados, D.; Mora-Garcia, S.; Cheng, J.C.; Nam, K.H.; Li, J.; Chory, J. BRL1 and BRL3 are novel brassinosteroid receptors that function in vascular differentiation in Arabidopsis. Development 2004 , 131 , 5341–5351. [ Google Scholar ] [ CrossRef ]
- Zhou, A.; Wang, H.; Walker, J.C.; Li, J. BRL1, a leucine-rich repeat receptor-like protein kinase, is functionally redundant with BRI1 in regulating Arabidopsis brassinosteroid signaling. Plant J. 2004 , 40 , 399–409. [ Google Scholar ] [ CrossRef ]
- Li, J.; Wen, J.; Lease, K.A.; Doke, J.T.; Tax, F.E.; Walker, J.C. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 2002 , 110 , 213–222. [ Google Scholar ] [ CrossRef ]
- Nam, K.H.; Li, J. BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 2002 , 110 , 203–212. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Hothorn, M.; Belkhadir, Y.; Dreux, M.; Dabi, T.; Noel, J.P.; Wilson, I.A.; Chory, J. Structural basis of steroid hormone perception by the receptor kinase BRI1. Nature 2011 , 474 , 467–471. [ Google Scholar ] [ CrossRef ]
- She, J.; Han, Z.; Kim, T.W.; Wang, J.; Cheng, W.; Chang, J.; Shi, S.; Wang, J.; Yang, M.; Wang, Z.Y.; et al. Structural insight into brassinosteroid perception by BRI1. Nature 2011 , 474 , 472–476. [ Google Scholar ] [ CrossRef ]
- Santiago, J.; Henzler, C.; Hothorn, M. Molecular mechanism for plant steroid receptor activation by somatic embryogenesis co-receptor kinases. Science 2013 , 341 , 889–892. [ Google Scholar ] [ CrossRef ]
- Wang, H.; Yang, C.; Zhang, C.; Wang, N.; Lu, D.; Wang, J.; Zhang, S.; Wang, Z.X.; Ma, H.; Wang, X. Dual role of BKI1 and 14-3-3s in brassinosteroid signaling to link receptor with transcription factors. Dev. Cell 2011 , 21 , 825–834. [ Google Scholar ] [ CrossRef ]
- Li, J.M.; Nam, K.H. Regulation of brassinosteroid signaling by a GSK3/SHAGGY-like kinase. Science 2002 , 295 , 1299–1301. [ Google Scholar ] [ CrossRef ]
- Wang, Z.Y.; Nakano, T.; Gendron, J.; He, J.; Chen, M.; Vafeados, D.; Yang, Y.; Fujioka, S.; Yoshida, S.; Asami, T.; et al. Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev. Cell 2002 , 2 , 505–513. [ Google Scholar ] [ CrossRef ]
- Yin, Y.; Wang, Z.Y.; Mora-Garcia, S.; Li, J.; Yoshida, S.; Asami, T.; Chory, J. BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 2002 , 109 , 181–191. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Tang, W.; Yuan, M.; Wang, R.; Yang, Y.; Wang, C.; Oses-Prieto, J.A.; Kim, T.W.; Zhou, H.W.; Deng, Z.; Gampala, S.S.; et al. PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat. Cell Biol. 2011 , 13 , 124–131. [ Google Scholar ] [ CrossRef ]
- Sun, Y.; Fan, X.Y.; Cao, D.M.; Tang, W.; He, K.; Zhu, J.Y.; He, J.X.; Bai, M.Y.; Zhu, S.; Oh, E.; et al. Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev. Cell 2010 , 19 , 765–777. [ Google Scholar ] [ CrossRef ]
- Yu, X.; Li, L.; Zola, J.; Aluru, M.; Ye, H.; Foudree, A.; Guo, H.; Anderson, S.; Aluru, S.; Liu, P.; et al. A brassinosteroid transcriptional network revealed by genome-wide identification of BES1 target genes in Arabidopsis thaliana . Plant J. 2011 , 65 , 634–646. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Nolan, T.M.; Vukasinovic, N.; Liu, D.; Russinova, E.; Yin, Y. Brassinosteroids: Multidimensional regulators of plant growth, development, and stress responses. Plant Cell 2020 , 32 , 295–318. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Lv, M.; Li, J. Molecular mechanisms of brassinosteroid-mediated responses to changing environments in Arabidopsis. Int. J. Mol. Sci. 2020 , 21 , 2737. [ Google Scholar ] [ CrossRef ]
- He, J.X.; Gendron, J.M.; Sun, Y.; Gampala, S.S.; Gendron, N.; Sun, C.Q.; Wang, Z.Y. BZR1 is a transcriptional repressor with dual roles in brassinosteroid homeostasis and growth responses. Science 2005 , 307 , 1634–1638. [ Google Scholar ] [ CrossRef ]
- Yin, Y.; Vafeados, D.; Tao, Y.; Yoshida, S.; Asami, T.; Chory, J. A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell 2005 , 120 , 249–259. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Gampala, S.S.; Kim, T.W.; He, J.X.; Tang, W.; Deng, Z.; Bai, M.Y.; Guan, S.; Lalonde, S.; Sun, Y.; Gendron, J.M.; et al. An essential role for 14-3-3 proteins in brassinosteroid signal transduction in Arabidopsis. Dev. Cell 2007 , 13 , 177–189. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Kim, T.W.; Guan, S.; Sun, Y.; Deng, Z.; Tang, W.; Shang, J.X.; Sun, Y.; Burlingame, A.L.; Wang, Z.Y. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nat. Cell Biol. 2009 , 11 , 1254–1260. [ Google Scholar ] [ CrossRef ]
- He, J.X.; Gendron, J.M.; Yang, Y.; Li, J.; Wang, Z.Y. The GSK3-like kinase BIN2 phosphorylates and destabilizes BZR1, a positive regulator of the brassinosteroid signaling pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2002 , 99 , 10185–10190. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Yang, M.; Li, C.; Cai, Z.; Hu, Y.; Nolan, T.; Yu, F.; Yin, Y.; Xie, Q.; Tang, G.; Wang, X. SINAT E3 ligases control the light-mediated stability of the brassinosteroid-activated transcription factor BES1 in Arabidopsis. Dev. Cell 2017 , 41 , 47–58.e4. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Kim, B.; Jeong, Y.J.; Corvalan, C.; Fujioka, S.; Cho, S.; Park, T.; Choe, S. Darkness and gulliver2/phyB mutation decrease the abundance of phosphorylated BZR1 to activate brassinosteroid signaling in Arabidopsis. Plant J. 2014 , 77 , 737–747. [ Google Scholar ] [ CrossRef ]
- Wang, Y.; Sun, S.; Zhu, W.; Jia, K.; Yang, H.; Wang, X. Strigolactone/MAX2-induced degradation of brassinosteroid transcriptional effector BES1 regulates shoot branching. Dev. Cell 2013 , 27 , 681–688. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Kim, E.J.; Lee, S.H.; Park, C.H.; Kim, S.H.; Hsu, C.C.; Xu, S.; Wang, Z.Y.; Kim, S.K.; Kim, T.W. Plant U-Box40 mediates degradation of the brassinosteroid-responsive transcription factor BZR1 in Arabidopsis roots. Plant Cell 2019 , 31 , 791–808. [ Google Scholar ] [ CrossRef ]
- Min, H.J.; Cui, L.H.; Oh, T.R.; Kim, J.H.; Kim, T.W.; Kim, W.T. OsBZR1 turnover mediated by OsSK22-regulated U-box E3 ligase OsPUB24 in rice BR response. Plant J. 2019 , 99 , 426–438. [ Google Scholar ] [ CrossRef ]
- Zheng, B.; Bai, Q.; Wu, L.; Liu, H.; Liu, Y.; Xu, W.; Li, G.; Ren, H.; She, X.; Wu, G. EMS1 and BRI1 control separate biological processes via extracellular domain diversity and intracellular domain conservation. Nat. Commun. 2019 , 10 , 4165. [ Google Scholar ] [ CrossRef ]
- Chen, W.; Lv, M.; Wang, Y.; Wang, P.A.; Cui, Y.; Li, M.; Wang, R.; Gou, X.; Li, J. BES1 is activated by EMS1-TPD1-SERK1/2-mediated signaling to control tapetum development in Arabidopsis thaliana . Nat. Commun. 2019 , 10 , 4164. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Cai, H.; Huang, Y.; Liu, L.; Zhang, M.; Chai, M.; Xi, X.; Aslam, M.; Wang, L.; Ma, S.; Su, H.; et al. Signaling by the EPFL-ERECTA family coordinates female germline specification through the BZR1 family in Arabidopsis. Plant Cell 2023 , 35 , 1455–1473. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Li, L.; Yu, X.; Thompson, A.; Guo, M.; Yoshida, S.; Asami, T.; Chory, J.; Yin, Y. Arabidopsis MYB30 is a direct target of BES1 and cooperates with BES1 to regulate brassinosteroid-induced gene expression. Plant J. 2009 , 58 , 275–286. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ye, H.; Li, L.; Guo, H.; Yin, Y. MYBL2 is a substrate of GSK3-like kinase BIN2 and acts as a corepressor of BES1 in brassinosteroid signaling pathway in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012 , 109 , 20142–20147. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, D.; Ye, H.; Guo, H.; Johnson, A.; Zhang, M.; Lin, H.; Yin, Y. Transcription factor HAT1 is phosphorylated by BIN2 kinase and mediates brassinosteroid repressed gene expression in Arabidopsis. Plant J. 2014 , 77 , 59–70. [ Google Scholar ] [ CrossRef ]
- Oh, E.; Zhu, J.Y.; Bai, M.Y.; Arenhart, R.A.; Sun, Y.; Wang, Z.Y. Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. Elife 2014 , 3 , e03031. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Yang, Z.; Yan, B.; Dong, H.; He, G.; Zhou, Y.; Sun, J. BIC1 acts as a transcriptional coactivator to promote brassinosteroid signaling and plant growth. EMBO J. 2021 , 40 , e104615. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Yang, R.; Liu, P.; Zhang, T.; Dong, H.; Jing, Y.; Yang, Z.; Tang, S.; Zhang, Y.; Lv, M.; Liu, J.; et al. Plant-specific BLISTER interacts with kinase BIN2 and BRASSINAZOLE RESISTANT1 during skotomorphogenesis. Plant Physiol. 2023 , 193 , 1580–1596. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Bai, M.Y.; Shang, J.X.; Oh, E.; Fan, M.; Bai, Y.; Zentella, R.; Sun, T.P.; Wang, Z.Y. Brassinosteroid, gibberellin and phytochrome impinge on a common transcription module in Arabidopsis. Nat. Cell Biol. 2012 , 14 , 810–817. [ Google Scholar ] [ CrossRef ]
- Plitsi, P.K.; Samakovli, D.; Roka, L.; Rampou, A.; Panagiotopoulos, K.; Koudounas, K.; Isaioglou, I.; Haralampidis, K.; Rigas, S.; Hatzopoulos, P.; et al. GA-mediated disruption of RGA/BZR1 complex requires HSP90 to promote hypocotyl elongation. Int. J. Mol. Sci. 2022 , 24 , 88. [ Google Scholar ] [ CrossRef ]
- Zhao, N.; Zhao, M.; Tian, Y.; Wang, Y.; Han, C.; Fan, M.; Guo, H.; Bai, M.Y. Interaction between BZR1 and EIN3 mediates signalling crosstalk between brassinosteroids and ethylene. New Phytol. 2021 , 232 , 2308–2323. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Wang, J.; Sun, N.; Zheng, L.; Zhang, F.; Xiang, M.; Chen, H.; Deng, X.W.; Wei, N. Brassinosteroids promote etiolated apical structures in darkness by amplifying the ethylene response via the EBF-EIN3/PIF3 circuit. Plant Cell 2023 , 35 , 390–408. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, J.; Chen, W.; Li, X.; Shi, H.; Lv, M.; He, L.; Bai, W.; Cheng, S.; Chu, J.; He, K.; et al. Jasmonates regulate apical hook development by repressing brassinosteroid biosynthesis and signaling. Plant Physiol. 2023 , 193 , 1561–1579. [ Google Scholar ] [ CrossRef ]
- Wang, L.; Tian, Y.; Shi, W.; Yu, P.; Hu, Y.; Lv, J.; Fu, C.; Fan, M.; Bai, M.Y. The miR396-GRFs module mediates the prevention of photo-oxidative damage by brassinosteroids during seedling de-etiolation in Arabidopsis. Plant Cell 2020 , 32 , 2525–2542. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Dong, H.; Liu, J.; He, G.; Liu, P.; Sun, J. Photoexcited phytochrome B interacts with brassinazole resistant 1 to repress brassinosteroid signaling in Arabidopsis. J. Integr. Plant Biol. 2020 , 62 , 652–667. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Wang, W.; Lu, X.; Li, L.; Lian, H.; Mao, Z.; Xu, P.; Guo, T.; Xu, F.; Du, S.; Cao, X.; et al. Photoexcited CRYPTOCHROME1 interacts with dephosphorylated BES1 to regulate brassinosteroid signaling and photomorphogenesis in Arabidopsis. Plant Cell 2018 , 30 , 1989–2005. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- He, G.; Liu, J.; Dong, H.; Sun, J. The blue-light receptor CRY1 interacts with BZR1 and BIN2 to modulate the phosphorylation and nuclear function of BZR1 in repressing BR signaling in Arabidopsis. Mol. Plant 2019 , 12 , 689–703. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Liang, T.; Mei, S.; Shi, C.; Yang, Y.; Peng, Y.; Ma, L.; Wang, F.; Li, X.; Huang, X.; Yin, Y.; et al. UVR8 interacts with BES1 and BIM1 to regulate transcription and photomorphogenesis in Arabidopsis. Dev. Cell 2018 , 44 , 512–523.e5. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ravindran, N.; Ramachandran, H.; Job, N.; Yadav, A.; Vaishak, K.P.; Datta, S. B-box protein BBX32 integrates light and brassinosteroid signals to inhibit cotyledon opening. Plant Physiol. 2021 , 187 , 446–461. [ Google Scholar ] [ CrossRef ]
- Li, M.; Li, P.; Wang, C.; Xu, H.; Wang, M.; Wang, Y.; Niu, X.; Xu, M.; Wang, H.; Qin, Y.; et al. Brassinosteroid signaling restricts root lignification by antagonizing SHORT-ROOT function in Arabidopsis. Plant Physiol. 2022 , 190 , 1182–1198. [ Google Scholar ] [ CrossRef ]
- Tian, Y.; Zhao, N.; Wang, M.; Zhou, W.; Guo, J.; Han, C.; Zhou, C.; Wang, W.; Wu, S.; Tang, W.; et al. Integrated regulation of periclinal cell division by transcriptional module of BZR1-SHR in Arabidopsis roots. New Phytol. 2022 , 233 , 795–808. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Vilarrasa-Blasi, J.; Gonzalez-Garcia, M.P.; Frigola, D.; Fabregas, N.; Alexiou, K.G.; Lopez-Bigas, N.; Rivas, S.; Jauneau, A.; Lohmann, J.U.; Benfey, P.N.; et al. Regulation of plant stem cell quiescence by a brassinosteroid signaling module. Dev. Cell 2014 , 30 , 36–47. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, T.; Xu, P.; Wang, W.; Wang, S.; Caruana, J.C.; Yang, H.Q.; Lian, H. Arabidopsis G-Protein beta subunit AGB1 interacts with BES1 to regulate brassinosteroid signaling and cell elongation. Front. Plant Sci. 2017 , 8 , 2225. [ Google Scholar ] [ CrossRef ]
- Zhang, D.; Jing, Y.; Jiang, Z.; Lin, R. The chromatin-remodeling factor PICKLE integrates brassinosteroid and gibberellin signaling during skotomorphogenic growth in Arabidopsis. Plant Cell 2014 , 26 , 2472–2485. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chai, G.; Qi, G.; Wang, D.; Zhuang, Y.; Xu, H.; Bai, Z.; Bai, M.Y.; Hu, R.; Wang, Z.Y.; Zhou, G.; et al. The CCCH zinc finger protein C3H15 negatively regulates cell elongation by inhibiting brassinosteroid signaling. Plant Physiol. 2022 , 189 , 285–300. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zhang, Y.; Li, B.; Xu, Y.; Li, H.; Li, S.; Zhang, D.; Mao, Z.; Guo, S.; Yang, C.; Weng, Y.; et al. The cyclophilin CYP20-2 modulates the conformation of BRASSINAZOLE-RESISTANT1, which binds the promoter of FLOWERING LOCUS D to regulate flowering in Arabidopsis. Plant Cell 2013 , 25 , 2504–2521. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ji, Y.; Qu, Y.; Jiang, Z.; Yan, J.; Chu, J.; Xu, M.; Su, X.; Yuan, H.; Wang, A. The mechanism for brassinosteroids suppressing climacteric fruit ripening. Plant Physiol. 2021 , 185 , 1875–1893. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Tian, X.; Li, X.; Zhou, W.; Ren, Y.; Wang, Z.; Liu, Z.; Tang, J.; Tong, H.; Fang, J.; Bu, Q. Transcription factor OsWRKY53 positively regulates brassinosteroid signaling and plant architecture. Plant Physiol. 2017 , 175 , 1337–1349. [ Google Scholar ] [ CrossRef ]
- Tian, X.; He, M.; Mei, E.; Zhang, B.; Tang, J.; Xu, M.; Liu, J.; Li, X.; Wang, Z.; Tang, W.; et al. WRKY53 integrates classic brassinosteroid signaling and the mitogen-activated protein kinase pathway to regulate rice architecture and seed size. Plant Cell 2021 , 33 , 2753–2775. [ Google Scholar ] [ CrossRef ]
- Ren, Y.; Tian, X.; Li, S.; Mei, E.; He, M.; Tang, J.; Xu, M.; Li, X.; Wang, Z.; Li, C.; et al. Oryza sativa mediator subunit OsMED25 interacts with OsBZR1 to regulate brassinosteroid signaling and plant architecture in rice. J. Integr. Plant Biol. 2020 , 62 , 793–811. [ Google Scholar ] [ CrossRef ]
- Gendreau, E.; Traas, J.; Desnos, T.; Grandjean, O.; Caboche, M.; Hofte, H. Cellular basis of hypocotyl growth in Arabidopsis thaliana . Plant Physiol. 1997 , 114 , 295–305. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chaiwanon, J.; Wang, W.; Zhu, J.Y.; Oh, E.; Wang, Z.Y. Information integration and communication in plant growth regulation. Cell 2016 , 164 , 1257–1268. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Su, D.; Xiang, W.; Liang, Q.; Wen, L.; Shi, Y.; Song, B.; Liu, Y.; Xian, Z.; Li, Z. Tomato SlBES1.8 influences leaf morphogenesis by mediating gibberellin metabolism and signaling. Plant Cell Physiol. 2022 , 63 , 535–549. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Fan, X.Y.; Sun, Y.; Cao, D.M.; Bai, M.Y.; Luo, X.M.; Yang, H.J.; Wei, C.Q.; Zhu, S.W.; Sun, Y.; Chong, K.; et al. BZS1, a B-box protein, promotes photomorphogenesis downstream of both brassinosteroid and light signaling pathways. Mol. Plant 2012 , 5 , 591–600. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Holtan, H.E.; Bandong, S.; Marion, C.M.; Adam, L.; Tiwari, S.; Shen, Y.; Maloof, J.N.; Maszle, D.R.; Ohto, M.; Preuss, S.; et al. BBX32, an Arabidopsis B-box protein, functions in light signaling by suppressing HY5-regulated gene expression and interacting with STH2/BBX21. Plant Physiol. 2011 , 156 , 2109–2123. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Ibanez, C.; Delker, C.; Martinez, C.; Burstenbinder, K.; Janitza, P.; Lippmann, R.; Ludwig, W.; Sun, H.; James, G.V.; Klecker, M.; et al. Brassinosteroids dominate hormonal regulation of plant thermomorphogenesis via BZR1. Curr. Biol. 2018 , 28 , 303–310.e3. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chaiwanon, J.; Wang, Z.Y. Spatiotemporal brassinosteroid signaling and antagonism with auxin pattern stem cell dynamics in Arabidopsis roots. Curr. Biol. 2015 , 25 , 1031–1042. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Sozzani, R.; Cui, H.; Moreno-Risueno, M.A.; Busch, W.; Van Norman, J.M.; Vernoux, T.; Brady, S.M.; Dewitte, W.; Murray, J.A.; Benfey, P.N. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 2010 , 466 , 128–132. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Zu, S.H.; Jiang, Y.T.; Chang, J.H.; Zhang, Y.J.; Xue, H.W.; Lin, W.H. Interaction of brassinosteroid and cytokinin promotes ovule initiation and increases seed number per silique in Arabidopsis. J. Integr. Plant Biol. 2022 , 64 , 702–716. [ Google Scholar ] [ CrossRef ]
- Chen, J.; Nolan, T.M.; Ye, H.; Zhang, M.; Tong, H.; Xin, P.; Chu, J.; Chu, C.; Li, Z.; Yin, Y. Arabidopsis WRKY46, WRKY54, and WRKY70 Transcription factors are involved in brassinosteroid-regulated plant growth and drought responses. Plant Cell 2017 , 29 , 1425–1439. [ Google Scholar ] [ CrossRef ]
- Ye, H.; Liu, S.; Tang, B.; Chen, J.; Xie, Z.; Nolan, T.M.; Jiang, H.; Guo, H.; Lin, H.Y.; Li, L.; et al. RD26 mediates crosstalk between drought and brassinosteroid signalling pathways. Nat. Commun. 2017 , 8 , 14573. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Xie, Z.; Nolan, T.; Jiang, H.; Tang, B.; Zhang, M.; Li, Z.; Yin, Y. The AP2/ERF transcription factor TINY modulates brassinosteroid-regulated plant growth and drought responses in Arabidopsis. Plant Cell 2019 , 31 , 1788–1806. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Tran, L.S.; Nakashima, K.; Sakuma, Y.; Simpson, S.D.; Fujita, Y.; Maruyama, K.; Fujita, M.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 2004 , 16 , 2481–2498. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Albertos, P.; Dundar, G.; Schenk, P.; Carrera, S.; Cavelius, P.; Sieberer, T.; Poppenberger, B. Transcription factor BES1 interacts with HSFA1 to promote heat stress resistance of plants. EMBO J. 2022 , 41 , e108664. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Rubin, G.; Tohge, T.; Matsuda, F.; Saito, K.; Scheible, W.R. Members of the LBD family of transcription factors repress anthocyanin synthesis and affect additional nitrogen responses in Arabidopsis. Plant Cell 2009 , 21 , 3567–3584. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Chai, S.; Chen, J.; Yue, X.; Li, C.; Zhang, Q.; de Dios, V.R.; Yao, Y.; Tan, W. Interaction of BES1 and LBD37 transcription factors modulates brassinosteroid-regulated root forging response under low nitrogen in arabidopsis. Front. Plant Sci. 2022 , 13 , 998961. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Loque, D.; von Wiren, N. Regulatory levels for the transport of ammonium in plant roots. J. Exp. Bot. 2004 , 55 , 1293–1305. [ Google Scholar ] [ CrossRef ]
- Suenaga, A.; Moriya, K.; Sonoda, Y.; Ikeda, A.; Von Wiren, N.; Hayakawa, T.; Yamaguchi, J.; Yamaya, T. Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants. Plant Cell Physiol. 2003 , 44 , 206–211. [ Google Scholar ] [ CrossRef ]
- Xuan, Y.H.; Priatama, R.A.; Huang, J.; Je, B.I.; Liu, J.M.; Park, S.J.; Piao, H.L.; Son, D.Y.; Lee, J.J.; Park, S.H.; et al. Indeterminate domain 10 regulates ammonium-mediated gene expression in rice roots. New Phytol. 2013 , 197 , 791–804. [ Google Scholar ] [ CrossRef ]
- Yang, S.; Yuan, D.; Zhang, Y.; Sun, Q.; Xuan, Y.H. BZR1 regulates brassinosteroid-mediated activation of AMT1;2 in rice. Front. Plant Sci. 2021 , 12 , 665883. [ Google Scholar ] [ CrossRef ]
- Jung, J.H.; Li, Z.; Chen, H.; Yang, S.; Li, D.; Priatama, R.A.; Kumar, V.; Xuan, Y.H. Mutation of phytochrome B promotes resistance to sheath blight and saline-alkaline stress via increasing ammonium uptake in rice. Plant J. 2023 , 113 , 277–290. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Liao, K.; Peng, Y.J.; Yuan, L.B.; Dai, Y.S.; Chen, Q.F.; Yu, L.J.; Bai, M.Y.; Zhang, W.Q.; Xie, L.J.; Xiao, S. Brassinosteroids antagonize jasmonate-activated plant defense responses through BRI1-EMS-SUPPRESSOR1 (BES1). Plant Physiol. 2020 , 182 , 1066–1082. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Qi, G.; Chen, H.; Wang, D.; Zheng, H.; Tang, X.; Guo, Z.; Cheng, J.; Chen, J.; Wang, Y.; Bai, M.Y.; et al. The BZR1-EDS1 module regulates plant growth-defense coordination. Mol Plant 2021 , 14 , 2072–2087. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Yuan, P.; Yang, S.; Feng, L.; Chu, J.; Dong, H.; Sun, J.; Chen, H.; Li, Z.; Yamamoto, N.; Zheng, A.; et al. Red-light receptor phytochrome B inhibits BZR1-NAC028-CAD8B signaling to negatively regulate rice resistance to sheath blight. Plant Cell Environ. 2023 , 46 , 1249–1263. [ Google Scholar ] [ CrossRef ] [ PubMed ]
- Xiao, S.; Hu, Q.; Zhang, X.; Si, H.; Liu, S.; Chen, L.; Chen, K.; Berne, S.; Yuan, D.; Lindsey, K.; et al. Orchestration of plant development and defense by indirect crosstalk of salicylic acid and brassinosteorid signaling via transcription factor GhTINY2. J. Exp. Bot. 2021 , 72 , 4721–4743. [ Google Scholar ] [ CrossRef ]
- Divi, U.K.; Krishna, P. Brassinosteroid: A biotechnological target for enhancing crop yield and stress tolerance. New Biotechnol. 2009 , 26 , 131–136. [ Google Scholar ] [ CrossRef ]
Click here to enlarge figure
| Gene Name | Locus | Effect on Plant Growth through Interaction with BES1/BZR1 | References |
|---|---|---|---|
| ARF6 | AT1G30330 | Promoting cell elongation and hypocotyl growth | [ ] |
| PIF4 | AT2G43010 | Promoting cell elongation and hypocotyl growth | [ ] |
| BIC1 | AT3G52740 | Promoting cell and hypocotyl elongation | [ ] |
| BLI | AT3G23980 | Promoting hypocotyl elongation in darkness | [ ] |
| RGA | AT2G01570 | Inhibiting cell elongation and BR-regulated plant growth | [ ] |
| HSP90 | AT5G52640 | Promoting hypocotyl elongation | [ ] |
| EIN3 | AT3G20770 | Promoting apical hook development | [ ] |
| SAUR17 | AT4G09530 | Promoting apical hook and closed cotyledon | [ ] |
| WAG2 | AT3G14370 | Inhibiting apical hook development | [ ] |
| GRF7 | AT5G53660 | Repressing chlorophyll biosynthesis promoting cell elongation | [ ] |
| phyB | AT2G18790 | Repressing BR signaling | [ ] |
| CRY1 | AT4G08920 | Inhibiting hypocotyl elongation under blue light | [ , ] |
| UVR8 | AT5G63860 | Repressing BR-regulated photomorphogenesis | [ ] |
| BBX32 | AT3G21150 | Inhibiting cotyledon opening | [ ] |
| SHR | AT4G37650 | Suppressing root lignification promoting periclinal division | [ , ] |
| BRAVO | AT5G17800 | Suppressing root quiescent center division | [ ] |
| AGB1 | AT4G34460 | Promoting cell elongation | [ ] |
| PKL | AT2G25170 | Promoting cell elongation | [ ] |
| C3H15 | AT1G68200 | Inhibiting cell elongation | [ ] |
| CYP20-2 | AT5G13120 | Promoting flowering | [ ] |
| ACO1 | AT2G19590 | Promoting fruit ripening | [ ] |
| OsWRKY53 | Os05g0343400 | Regulating rice architecture and increasing seed size | [ , ] |
| OsMED25 | Os09g0306700 | Regulating rice architecture | [ ] |
| Gene Name | Locus | Effect on Stress Tolerance through Interaction with BES1/BZR1 | References |
|---|---|---|---|
| WRKY46 | AT2G46400 | Suppressing drought response | [ ] |
| WRKY54 | AT2G40750 | Suppressing drought response | |
| WRKY70 | AT3G56400 | Suppressing drought response | |
| RD26 | AT4G27410 | Inhibiting BR-regulated growth under drought condition | [ ] |
| TINY | AT5G25810 | Inhibiting BR-regulated growth under drought condition | [ ] |
| HSFA1a | AT4G17750 | Improving heat stress | [ ] |
| LBD37 | AT5G67420 | Promoting nitrogen response | [ ] |
| OsIDD10 | Os04g47860 | Improving nitrogen uptake | [ ] |
| phyB | AT2G18790 | Inhibiting nitrogen uptake and sheath blight resistance | [ , ] |
| MYB34 | AT5G60890 | Suppressing insect defense | [ ] |
| MYB122 | AT1G74080 | Suppressing insect defense | [ ] |
| MYB51 | AT1G18570 | Suppressing insect defense | [ ] |
| EDS1 | AT3G48090 | Increasing pathogen resistance | [ ] |
| GhTINY2 | GhD06G0642 | Increased immune response | [ ] |
| The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
Share and Cite
Cao, X.; Wei, Y.; Shen, B.; Liu, L.; Mao, J. Interaction of the Transcription Factors BES1/BZR1 in Plant Growth and Stress Response. Int. J. Mol. Sci. 2024 , 25 , 6836. https://doi.org/10.3390/ijms25136836
Cao X, Wei Y, Shen B, Liu L, Mao J. Interaction of the Transcription Factors BES1/BZR1 in Plant Growth and Stress Response. International Journal of Molecular Sciences . 2024; 25(13):6836. https://doi.org/10.3390/ijms25136836
Cao, Xuehua, Yanni Wei, Biaodi Shen, Linchuan Liu, and Juan Mao. 2024. "Interaction of the Transcription Factors BES1/BZR1 in Plant Growth and Stress Response" International Journal of Molecular Sciences 25, no. 13: 6836. https://doi.org/10.3390/ijms25136836
Article Metrics
Article access statistics, further information, mdpi initiatives, follow mdpi.

Subscribe to receive issue release notifications and newsletters from MDPI journals
Lab on a Chip
An integrative temperature-controlled microfluidic system for budding yeast heat shock response analysis in single cell level.
Cells can respond and adapt to complex forms of environmental change. Budding yeast is wildly used as a model system for these stress response studies. In these studies, the precise control of the environment with high temporal resolution is most important. However, there is a lack of single-cell research platforms that enable precise control of the temperature and form of cell growth. This has hindered our understanding of cellular coping strategies in the face of diverse forms of temperature change. Here, we developed a novel temperature-controlled microfluidic platform that integrates a micro-heater(using liquid metal) and thermocouple(liquid metal vs conductive PDMS) on a chip. Three forms of temperature changes: step, gradient, and periodical oscillations were realized by automated equipment. The platform has the advantages of low cost and a simple fabrication process. Moreover, we investigated the nuclear entry and exit behaviors of the transcription factor Msn2 in yeast in response to heat stress (37°C) with different heating modes. The feasibility of this temperature-controlled platform for studying the protein dynamic behavior of yeast cells was demonstrated.
Supplementary files
- Supplementary information PDF (1027K)
Article information
Download citation, permissions.
J. Hong, H. He, Y. Xu, S. Wang and C. Luo, Lab Chip , 2024, Accepted Manuscript , DOI: 10.1039/D4LC00313F
To request permission to reproduce material from this article, please go to the Copyright Clearance Center request page .
If you are an author contributing to an RSC publication, you do not need to request permission provided correct acknowledgement is given.
If you are the author of this article, you do not need to request permission to reproduce figures and diagrams provided correct acknowledgement is given. If you want to reproduce the whole article in a third-party publication (excluding your thesis/dissertation for which permission is not required) please go to the Copyright Clearance Center request page .
Read more about how to correctly acknowledge RSC content .
Social activity
Search articles by author.
This article has not yet been cited.
Advertisements
News : The 2024 HST graduation

Photo credit: Justin Knight
There were 57 clinician-scientists in this year’s graduating class, 40 attended the ceremony
Mindy Blodgett | IMES-HST
The 2024 graduating class of the Harvard-MIT Program in Health Sciences and Technology (HST) gathered on May 22, to celebrate their accomplishments with their families and friends, at the MIT Bartos Theater & Atrium. Also in attendance were HST alumni, faculty, and staff.
This 2024 graduation class includes 57 graduates: 35 MD graduates, and 25 Medical Engineering and Medical Physics (MEMP)PhDs; one Master of Science graduate, and one Graduate Education in Medical Sciences, or GEMS certificate, recipient. There were 40 graduates in attendance. HST MD graduates also participated in Harvard graduation events on May 23, and graduates of the HST Medical Engineering and Medical Physics (MEMP) PhD program participated in the MIT School of Engineering Advanced Degree Ceremony, and hooding event, on May 29.

All enjoyed congratulatory remarks from HST Associate Director Richard N. Mitchell, MD, PhD; Dean of the Harvard Medical School (HMS) George Q. Daley, MD (HST ’91), PhD; and Elazer Edelman, MD (HST ’83), PhD (HST ’84), Director of the Institute for Medical Engineering and Science (IMES). Also participating in the ceremony were Wolfram Goessling MD, PhD, the co-director of HST at Harvard, Collin M. Stultz, MD, (HST ’97), PhD, co-director of HST at MIT, and associate director of IMES (IMES is HST’s home at MIT), as well as Junne Kamihara, Associate Director, MD Advising, HST, and HST Associate Director, Matthew Frosch.

L to R, Junne Kamihara, Associate Director, MD Advising, HST; HST Associate Director Richard N. Mitchell, MD, PhD. Photo credit: Justin Knight
Dean Daley, an HST alumnus, called the occasion, a “spectacular achievement to graduate from the country’s pre-eminent program in translational biomedical science and engineering” and he praised the graduates’ “persistence in getting through the pandemic,” as Covid was at its height when many from the class began their studies in 2020. Daley observed that the graduates will witness “explosive developments” during their careers, in such areas as gene editing, artificial intelligence (AI) and the needs of an aging population.

Harvard Medical School (HMS) George Q. Daley, MD (HST ’91), PhD. Photo credit: Justin Knight.
Stultz called addressing the graduates “one of the best parts of my job,” remarking that “few individuals have achieved your level of accomplishments.”

Collin M. Stultz, MD, (HST ’97), PhD, co-director of HST at MIT, and associate director of IMES. Photo credit: Justin Knight.

Wolfram Goessling MD, PhD, the co-director of HST at Harvard, congratulated the graduates and their families and friends. Photo credit: Justin Knight.
Edelman, an HST alumnus, who is also a senior attending physician, Brigham and Women’s Hospital, shared a story about one of his patients, a middle school principal from Western Massachusetts, who was the “heart and soul” of his school, and of his small town. He said that the graduates were chosen for HST because “of what we saw in you…your heart and soul” and that “together, we can harness medicine to make the world a better place.”

Elazer Edelman, MD (HST ’83), PhD (HST ’84), Director of the Institute for Medical Engineering and Science (IMES). Photo credit: Justin Knight.
Abby Aymond, an HST MD graduate, was the 2024 class speaker. She praised the “exceptional sense of community and friendship” she had experienced while a student at HST. She said the some of the lessons she was taking from her years at HST were to “relax all the noise…focus only on the problem at hand…and to always be open to new information.”

Abby Aymond, HST MD graduate, was the 2024 class speaker. Photo credit: Justin Knight.

Elazer Edelman, left, and George Daley, right, address the graduates at the end of the ceremony, urging them to stay in touch. Photo credit: Justin Knight.

HST Associate Director Richard N. Mitchell donned the traditional Red Sox graduation cap, and applauded the graduates. Photo credit: Justin Knight.
The HST 2024 Graduates:
Doctor of Medicine
Medical Sciences
Abby Aymond, BS
Thesis Topic: Optimization of Ventricular Efficiency and Renal Artery Perfusion in a Bench Top Model System
Alaleh Azhir, BS
Thesis Topic: Chromosomes vs Hormones: Decoding the Expression Mosaic in Liver and Adipose Tissues
James Diao, BS
summa cum laude
The Seidman Prize for Outstanding HST Senior Medical Student Thesis
Richard C. Cabot Prize
Thesis Topic: The Use of Race in Clinical Algorithms
Christopher Michael Dietrich, BS
Thesis Topic: Towards Treat-Seq: Predicting Therapeutic Response from Transcriptomic Signatures
Jonah Issac Donnenfield, BA
magna cum laude
Thesis Topic: Transcriptomic Profiling of the Post-traumatic Porcine Knee: Degenerative Pathophysiology and Machine Learning Application
Micayla Flores, SB
Thesis Topic: Ambulatory and Delivery Obstetric Comorbidity Index (OB-CMI) for Identification of Pregnant Individuals at Risk for Severe Maternal Morbidity (SMM)
Allyson Freedy, BA, PhD
Leon Reznick Memorial Prize
HMS Multiculturalism Award
Thesis Topic: Uncovering the Biology of Chromatin Regulators with Drug Resistance Alleles
William Hao Ge, BS
Thesis Topic: Stereotypic Patterns and Genomic Correlates of Organotropism in Metastatic Melanoma
Blake Hauser, BSPH, PhD
Thesis Topic: Structure-Based Network Analysis Predicts Pathogenic Variants in Human Proteins Associated with Inherited Retinal Disease
Sofia Hu, BA, PhD
Thesis Topic: Transcription Factor Antagonism Regulates Heterogeneity in Embryonic Stem Cell States
Nauman Javed, BS, PhD
Thesis Topic: Strategies for Characterizing the Regulatory Code of the Human Genome
Tushar Vinod Kamath, SB, SM, PhD
Thesis Topic: Cell States and Neuronal Vulnerabilities in Neurodegenerative Diseases
Minjee Kim, BA
Thesis Topic: Transcriptional Antagonism by CDK8 Inhibition Improves Therapeutic Efficacy of MEK Inhibitors
Patrick Lenehan, BS, PhD
Thesis Topic: Investigating the Impact of Eosinophils on Pancreatic Cancer Growth and Metastasis
Claudio Macias Trevino, BS, PhD
Thesis Topic: Transcriptional Regulation of Esrp1 and its Role in Craniofacial Morphogenesis
Eliana Marostica, BA, MBMI
Thesis Topic: Systematic Quantification of Morphological Patterns in Surgical Specimens of Cancers
Eduardo Maury, SB, PhD
Thesis Topic: Somatic Mutations in the Human Brain: Tracing the Origins of Cancer and Schizophrenia
Elizabeth Minten, BS, PhD
Thesis Topic: Role of CDK12 in R-Loop Formation
Katherine Nabel Smith, BS, PhD
Thesis Topic: Molecular Mechanisms for Broad Neutralization of Emerging RNA Viruses
Julia E. Page, SB, PhD
Thesis Topic: Peptidoglycan Hydrolases, their Protein Partners, and Related Membrane Proteins in Staphylococcus Aureus
Deborah Plana, SB, PhD
Thesis Topic: Clinical Trial Data Science to Advance Precision Oncology
Sheridan Rea, BS, MS
Thesis Topic: Retrospective Cohort Analysis of Sociodemographic Factors and Postpartum Hemorrhage Outcomes
Sara Ann Rubin, BA, PhD
Thesis Topic: Zebrafish Immune Cell Development and Diversity in Health and Disease
Jamie Erin Shade, BS
Thesis Topic: Relationships Between Cardiac Magnetic Resonance-derived Myocardial, Hepatic, and Splenic Extracellular Volumes in Patients after the Fontan Operation
Bryce Filip Starr, BS
Thesis Topic: Generation and Validation of a Bileaflet Venous Valve for Single Ventricle Physiology
Hannah Jacqueline Szapary, BS, SM
Thesis Topic: Mechanical and Biologic Impact of Dynamic Loading on Bovine and Human Models of Osteoarthritis
Max Louis Valenstein, BS, MS, PhD
Thesis Topic: Integration of Amino Acid Signals by the mTORC1 Pathway
Sarah Weiss, SB, PhD
Thesis Topic: Deletion of an Exhaustion-specific PD-1 Enhancer Modulates CD8+ T Cell Fate and Function
Omar Yaghi, BS, PhD
Thesis Topic: Uncovering Stromal Cell Functions in Acute and Chronic Muscle Injuries
Katherine Young, SB, MEng
Thesis Topic: Transmission and Evolution of Staphylococcus Aureus in Families with Atopic Dermatitis
Doctor of Philosophy
Medical Engineering/Medical Physics
Jon Arizti Sanz, MNG
Thesis Topic: From Sample to Answer: Innovations in Sample Processing and CRISPR-based Diagnostics for Enhanced Clinical Translation and Field Deployment
Olivia Jane Arnold, SB
Thesis Topic: Therapeutic Applications of DNA Origami-based Progammable Nanoparticles
Rachel Bellisle, SB
Thesis Topic: A Wearable Countermeasure for Musculoskeletal Deconditioning in Human Spaceflight
Adam G. Berger, SB
Thesis Topic: Systematic Engineering of Controlled, Localized Oligonucleotide Delivery Systems for Wound Angiogenesis
Jennifer Dawkins, SB
Thesis Topic: Computational Prediction of Health Status from the Human Gut Microbiome and Metabolome
Brian Tshao Do, SB
Thesis Topic: Metabolic and Genetic Factors Guiding Hematopoietic Cell Fate
Mingjian He, SB
Thesis Topic: State-space Modeling of Neural Oscillations: Toward Assessing Alzheimer’s Disease Neuropathology with Sleep EEG
Brennan Leo Jackson, SB
Thesis Topic: The Impact of Gamma Stimulation on Neurological Phenotypes of Alzheimer's Dementia and Down Syndrome
Morgan Elizabeth Janes, SB
Thesis Topic: Engineering Translational Vaccine Delivery Systems with the Polyphenol Tannic Acid
Ashwin Srinivasan Kumar, BNG
Thesis Topic: Targeting B Cells to Improve Therapeutic Outcomes for Pediatric Medulloblastoma
Christian Landeros, SB
Thesis Topic: Machine-Guided Biopsy Analysis in Oncology: Facilitating Diagnostic Access and Biomedical Discovery Through Deep Learning
Ben D. Leaker, BNG
Thesis Topic: Biological and Biomechanical Effects of Direct Perturbation of Tissue Structure in the Cirrhotic Liver
Fiona Macleod, BNG
Thesis Topic: Investigating the Fidelity of Classic Cardiovascular Metrics in the Context of a Failing and Mechanically Supported Heart
Maria Carmen Martin Alonso, MNG
Thesis Topic: Amplifying Signals in the Tumor Microenvironment for Drug Development and Diagnostics
Eli Mattingly, SB
Thesis Topic: Design, Construction, and Validation of Magnetic Particle Imaging Systems for Rodent, Primate, and Human Functional Neuroimaging
Vincent Miao, BNG
Thesis Topic: Profiling Host Respiratory Responses to SARS-CoV-2 Infection
Allison Paige Porter, SB
Thesis Topic: Automation Framework for Exploration Medicine (AFEM): A Path for Integrating Automation into Autonomous Emergency Care
Rumya Raghavan, SB
Thesis Topic: Engineering Minimally Immunogenic Cargos and Delivery Modalities for Gene Therapy
Michelle Ramseier, SB
Thesis Topic: Cooptation of B Cell Developmental States in Malignancy and Autoimmunity
Luca Rosalia, MNG
Thesis Topic: Soft Robotic Platforms for the Simulation of Cardiovascular Disease and Device Development
Daphne Schlesinger, SB
Thesis Topic: Physiology-Inspired Deep Learning for Improved Heart Failure Management
Sydney Sherman, SB
Thesis Topic: Single-sided Magnetic Resonance Sensors for Clinical Detection of Volume Status
Nalini Singh, SB
Thesis Topic: Physics-Inspired Deep Learning for Inverse Problems in MRI
Anubhav Sinha, SB, MNG
Thesis Topic: Spatially Precise in situ Transcriptomics in Intact Biological Systems
Mingyu Yang, SB
Thesis Topic: Myelination Diseases of the Central Nervous System: Artificial Axons as in Vitro Models of Chemomechanical Cues
Master of Science
Health Sciences and Technology
Noah Stanley Warner, SB
Thesis Topic: A Framework for Detection and Observation of Radiation Chemistry Species on an MR-Linac
Certificate
Graduate Education in Medical Sciences
Akshay Kothakonda, BNG, SM
Thesis Topic: Engineering Mechanical Counter Pressure Spacesuits and Compression Garments: Active Pressurization and Design for Mobility

IMAGES
VIDEO
COMMENTS
computational approaches for transcription factor specificity engineering to expand the breadth of potential biosensor applications, and creating chimeric transcription factors to expedite biosensor discovery and overcome the currently high levels of characterization typically required to develop biosensors.
1. Introduction. Transcription is a crucial component of the central dogma of molecular biology (DNA-RNA-protein) [], serving as a bridge that translates genetic information into diverse forms at the individual level.Transcription factors (TFs) play a pivotal role in regulating the transcription of target genes by selectively recognizing and binding specific DNA regions known as TF binding ...
Transcription factors are proteins that initiate and modulate transcription rate by interacting with specific DNA recognition sequences in the target genes. As shown in Fig. 1, these DNA-binding transcription factors are structurally classified into four major classes: Helix-turn-helix homeodomain (e.g. PBX1 ), C. 2. H. 2 . zinc
Investigating the role of transcription factor, Trl, during germline development in the Drosophila ovary by Lindsay L. Davenport July 2019 Director of Thesis: Elizabeth T. Ables, Ph. D. Major Department: Biology Oogenesis is the process by which an egg develops from undifferentiated cells in the ovary.
scription factors. In this thesis, we focus on the binding of transcription factors to upstream region motifs to understand the mechanism of gene regulation. Sonic hedgehog (Shh) signals direct digit number and identity in the vertebrate limb via Gli transcription factors. We sought to identify key Gli binding motifs in
Introduction. The sine oculis (SIX) homeobox family of transcription factors play important developmental roles in a wide range of species from fruit flies to humans. The founding member, sine oculis (so), was first identified in Drosophila melanogaster where it was discovered to be required for compound eye formation (Cheyette et al., 1994; Serikaku and O'Tousa, 1994).
PREDICTING TRANSCRIPTION FACTOR BINDING USING NEURAL STRUCTURED LEARNING A Thesis in Bioinformatics and Genomics by Natalie Zesati Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science December 2020. ii The thesis of Natalie Zesati was reviewed and approved by the following: Shaun Mahony Assistant Professor of ...
I have also applied supervised learning methods for predicting transcription factor binding locations based on combinations of regulatory motifs. For each experiment in a compendium of ChIP-chip studies, I constructed a classifier to distinguish between regions bound by the given factor and regions bound by any other factor. For each
Transcription factors (TFs) transcriptionally regulate genes by binding nearby sequence elements. The evolutionary mechanisms driving the evolution of TF binding events between species are unclear. This thesis addresses three disparate predictions of natural selection acting on different
Transcription factors (TFs) recognize specific DNA sequences to control chromatin and transcription, forming a complex system that guides expression of the genome. Despite keen interest in understanding how TFs control gene expression, it remains challenging to determine how the precise genomic binding sites of TFs are specified and how TF ...
Background: Transcriptional factors (TFs) are responsible for regulating the transcription of pro-oncogenes and tumor suppressor genes in the process of tumor development. However, the role of these transcription factors in Bladder cancer (BCa) remains unclear. And the main purpose of this research is to explore the possibility of these TFs serving as biomarkers for BCa.
Thesis proposal Functional Validation of Transcription Factor to Gene Interactions by Statistical Learning of Gaussian Bayesian networks from SNP and Expression data. Jing Xiang Machine Learning Department Carnegie Mellon University [email protected] Committee members: Seyoung Kim Geoff Gordon Carl Kingsford Steffi Oesterreich January 23, 2017
This thesis explores the role of transcription factors in sensory neuron specification. We describe the transcription factor Foxs1 as an early sensory neuronal marker and use it to
Hinojosa, Leetoria, Investigating the Localization of FOXO Transcription Factors in. Glioblastoma. Master of Sciences (MS), May, 2020, 32pp., 1 table, 7 figures, 17 references. The Phosphatidylinositol 3 Kinase (PI3K) pathway is an essential intracellular signaling. pathway that regulates cellular growth, survival, and fate.
Transcription Factors. Transcription factors are proteins that bind to DNA-regulatory sequences (enhancers and silencers), usually localized in the 5 -upstream region of target genes, to modulate the rate of gene transcription. This may result in increased or decreased gene transcription, protein synthesis, and subsequent altered cellular function.
Illustration of an activator. In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription of genetic information from DNA to messenger RNA, by binding to a specific DNA sequence. The function of TFs is to regulate—turn on and off—genes in order to make sure that they are expressed in the desired cells at ...
Transcription factors are proteins that help turn specific genes "on" or "off" by binding to nearby DNA. Transcription factors that are activators boost a gene's transcription. Repressors decrease transcription. Groups of transcription factor binding sites called enhancers and silencers can turn a gene on/off in specific parts of the body.
a Summary statistics of transcription factor (TF)-target gene(TG) links. b Peak annotation. c137 Distributions of number of co-binding TF), TGs, and peaks for individual transcription factors. 138 d Correlations between the numbers of co-binding transcription factors and target genes and the numbers 139 of co-binding TFs and the number of peaks.
Fullscreen. Transcription factor binding sites identification using machine learning techniques. Cite. Download(3.41 MB) Embed. thesis. posted on2023-01-18, 15:54authored byHai Thanh Do. Submission note: A thesis submitted in total fulfilment of the requirements for the degree of Doctor of Philosophy to the Department of Computer Science and ...
Here, we isolated and functionally characterized SmERF1L1, a novel JA (Jasmonic acid)-responsive gene encoding AP2/ERF transcription factor, from Salvia miltiorrhiza. SmERF1L1 was responsive to methyl jasmonate (MJ), yeast extraction (YE), salicylic acid (SA) and ethylene treatments. Subcellular localization assay indicated that SmERF1L1 ...
The plant-specific YABBY transcription factor family plays important roles in plant growth and development, particularly leaf growth, floral organ formation, and secondary metabolite synthesis. Here, we identified a total of 13 OfYABBY genes from the Osmanthus fragrans genome. These 13 OfYABBY genes were divided into five subfamilies through phylogenetic analysis, and genes in the same ...
We have investigated the contact points of a positive transcription factor with the internal control region of the 5S ribosomal RNA genes of Xenopus. The methylation of any one of eight G residues clustered at the 3′ end of the internal control region on the noncoding strand of the DNA or the ethylation of their surrounding phosphates interferes with the binding of this protein.
1. Introduction. Transcription factors (TFs) are a group of mediators that bind the promoter or regulatory sequence of a gene to control its rate of transcribing genetic information from DNA to messenger RNA ().This transcription control is key to ensuring an adequate level of expression of a given protein in targeted cells at a particular developmental stage.
A high-quality single-cell chromatin accessibility atlas of colorectal cancer epithelial cells identified two epigenetic subgroups that match intrinsic-consensus molecular subtypes along with key transcription factors and their synergistic modules that regulate subtype-specific phenotypic features.
The mitochondrial transcription factor A, TFAM, has a dual function in the organelle: it activates mitochondrial DNA transcription by binding to the HSP and LSP promoters, while in higher concentrations compacts the mtDNA. In this thesis the mechanism of complex formation between the mitochondrial transcription factor A
The binding of CaHDZ15 to the promoter of CaHSFA6a and thus the transcription of CaHSFA6a were enhanced by the presence of CaHsp70-2 , indicating that CaHsp70-2 contributes to thermotolerance at least partially by acting as a transcriptional coactivator for CaHDZ15 to activate the transcription of CaHSFA6a, similar to HSP70-14 which interacts ...
TRANSCRIPTION FACTOR BINDING SITES By LIANG ZHAO Bachelor of Science Zhejiang University Hangzhou, China 1992 . . Master of Engineering BetJmg Research Institute of Chemical Industry Beijing, China 1995 Submitted to the Faculty of the Graduate College of the Oklahoma State University in partial fulfillment of the requirements for the degree of
Bri1-EMS Suppressor 1 (BES1) and Brassinazole Resistant 1 (BZR1) are two key transcription factors in the brassinosteroid (BR) signaling pathway, serving as crucial integrators that connect various signaling pathways in plants. Extensive genetic and biochemical studies have revealed that BES1 and BZR1, along with other protein factors, form a complex interaction network that governs plant ...
Moreover, we investigated the nuclear entry and exit behaviors of the transcription factor Msn2 in yeast in response to heat stress (37°C) with different heating modes. The feasibility of this temperature-controlled platform for studying the protein dynamic behavior of yeast cells was demonstrated.
Thesis Topic: Structure-Based Network Analysis Predicts Pathogenic Variants in Human Proteins Associated with Inherited Retinal Disease. Sofia Hu, BA, PhD. Thesis Topic: Transcription Factor Antagonism Regulates Heterogeneity in Embryonic Stem Cell States. Nauman Javed, BS, PhD